Difference between revisions of "YitJ"
| Line 118: | Line 118: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 894 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
Revision as of 10:04, 17 April 2014
- Description: methionine synthase
| Gene name | yitJ |
| Synonyms | |
| Essential | no |
| Product | methionine synthase |
| Function | methionine biosynthesis |
| Gene expression levels in SubtiExpress: yitJ | |
| Metabolic function and regulation of this protein in SubtiPathways: yitJ | |
| MW, pI | 67 kDa, 5.336 |
| Gene length, protein length | 1836 bp, 612 aa |
| Immediate neighbours | yitI, yitK |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 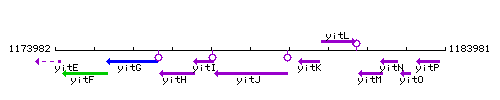
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed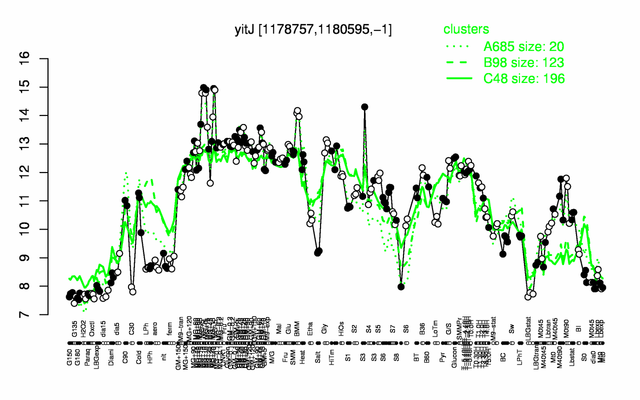
| |
Contents
Categories containing this gene/protein
membrane proteins, biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU11010
Phenotypes of a mutant
Database entries
- BsubCyc: BSU11010
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- membrane associated PubMed
Database entries
- BsubCyc: BSU11010
- Structure:
- UniProt: O06745
- KEGG entry: [2]
- E.C. number: 2.1.1.13
Additional information
Expression and regulation
- Sigma factor:
- Regulatory mechanism: S-box: transcription termination/ antitermination, the S-box riboswitch binds S-adenosylmethionine resulting in termination PubMed
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 894 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
The S-box
Vamsi Krishna Boyapati, Wei Huang, Jessica Spedale, Fareed Aboul-Ela
Basis for ligand discrimination between ON and OFF state riboswitch conformations: the case of the SAM-I riboswitch.
RNA: 2012, 18(6);1230-43
[PubMed:22543867]
[WorldCat.org]
[DOI]
(I p)
Changrui Lu, Fang Ding, Anirban Chowdhury, Vineeta Pradhan, Jerneja Tomsic, W Michael Holmes, Tina M Henkin, Ailong Ke
SAM recognition and conformational switching mechanism in the Bacillus subtilis yitJ S box/SAM-I riboswitch.
J Mol Biol: 2010, 404(5);803-18
[PubMed:20951706]
[WorldCat.org]
[DOI]
(I p)
Ana Gutiérrez-Preciado, Tina M Henkin, Frank J Grundy, Charles Yanofsky, Enrique Merino
Biochemical features and functional implications of the RNA-based T-box regulatory mechanism.
Microbiol Mol Biol Rev: 2009, 73(1);36-61
[PubMed:19258532]
[WorldCat.org]
[DOI]
(I p)
Jerneja Tomsic, Brooke A McDaniel, Frank J Grundy, Tina M Henkin
Natural variability in S-adenosylmethionine (SAM)-dependent riboswitches: S-box elements in bacillus subtilis exhibit differential sensitivity to SAM In vivo and in vitro.
J Bacteriol: 2008, 190(3);823-33
[PubMed:18039762]
[WorldCat.org]
[DOI]
(I p)
F J Grundy, T M Henkin
The S box regulon: a new global transcription termination control system for methionine and cysteine biosynthesis genes in gram-positive bacteria.
Mol Microbiol: 1998, 30(4);737-49
[PubMed:10094622]
[WorldCat.org]
[DOI]
(P p)
Other publications