Difference between revisions of "YesZ"
Raphael2215 (talk | contribs) |
|||
| Line 13: | Line 13: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || unknown | |style="background:#ABCDEF;" align="center"|'''Function''' || unknown | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU07080 yesZ] |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 73 kDa, 5.5 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 73 kDa, 5.5 | ||
| Line 21: | Line 21: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yesY]]'', ''[[yetA]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yesY]]'', ''[[yetA]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU07080 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU07080 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU07080 Advanced_DNA] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:yesZ_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:yesZ_context.gif]] | ||
Revision as of 12:19, 13 May 2013
- Description: beta-galactosidase
| Gene name | yesZ |
| Synonyms | |
| Essential | no |
| Product | beta-galactosidase |
| Function | unknown |
| Gene expression levels in SubtiExpress: yesZ | |
| MW, pI | 73 kDa, 5.5 |
| Gene length, protein length | 1989 bp, 663 aa |
| Immediate neighbours | yesY, yetA |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 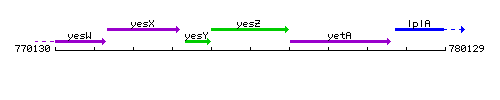
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed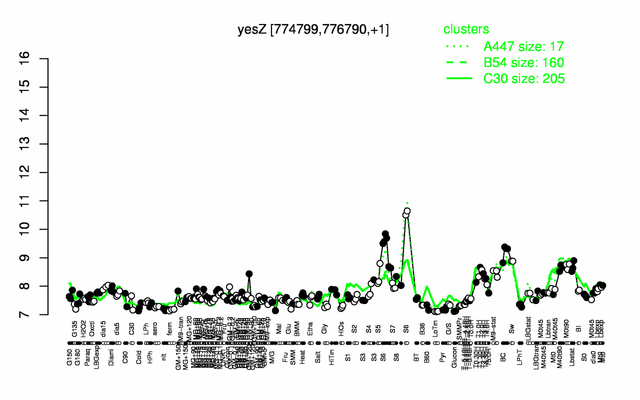
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU07080
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Hydrolysis of terminal non-reducing beta-D-galactose residues in beta-D-galactosides (according to Swiss-Prot)
- Protein family: glycosyl hydrolase 42 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O31529
- KEGG entry: [2]
- E.C. number: 3.2.1.23
Additional information
Expression and regulation
- Sigma factor:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Sandrine Poncet, Maryline Soret, Peggy Mervelet, Josef Deutscher, Philippe Noirot
Transcriptional activator YesS is stimulated by histidine-phosphorylated HPr of the Bacillus subtilis phosphotransferase system.
J Biol Chem: 2009, 284(41);28188-28197
[PubMed:19651770]
[WorldCat.org]
[DOI]
(I p)
Akihito Ochiai, Takafumi Itoh, Akiko Kawamata, Wataru Hashimoto, Kousaku Murata
Plant cell wall degradation by saprophytic Bacillus subtilis strains: gene clusters responsible for rhamnogalacturonan depolymerization.
Appl Environ Microbiol: 2007, 73(12);3803-13
[PubMed:17449691]
[WorldCat.org]
[DOI]
(P p)
Stephanie Shipkowski, Jean E Brenchley
Bioinformatic, genetic, and biochemical evidence that some glycoside hydrolase family 42 beta-galactosidases are arabinogalactan type I oligomer hydrolases.
Appl Environ Microbiol: 2006, 72(12);7730-8
[PubMed:17056685]
[WorldCat.org]
[DOI]
(P p)