Difference between revisions of "YesW"
(→References) |
|||
| Line 125: | Line 125: | ||
=References= | =References= | ||
| − | <pubmed> 19651770 17449691,17947240 , 19193638 </pubmed> | + | <pubmed> 19651770 17449691,17947240 , 19193638 16682770 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:06, 26 December 2010
- Description: rhamnogalacturonan lyase, generates oligosaccharides
| Gene name | yesW |
| Synonyms | |
| Essential | no |
| Product | rhamnogalacturonan lyase, generates oligosaccharides |
| Function | utilization of rhamnogalacturonan |
| MW, pI | 67 kDa, 5.966 |
| Gene length, protein length | 1860 bp, 620 aa |
| Immediate neighbours | yesV, yesX |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 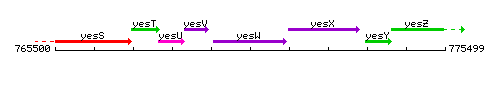
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU07050
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: FG-GAP repeat (according to Swiss-Prot)
- Paralogous protein(s): YesX
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: secreted (according to Swiss-Prot)
Database entries
- Structure: 2Z8R
- UniProt: O31526
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Sandrine Poncet, Maryline Soret, Peggy Mervelet, Josef Deutscher, Philippe Noirot
Transcriptional activator YesS is stimulated by histidine-phosphorylated HPr of the Bacillus subtilis phosphotransferase system.
J Biol Chem: 2009, 284(41);28188-28197
[PubMed:19651770]
[WorldCat.org]
[DOI]
(I p)
Akihito Ochiai, Takafumi Itoh, Bunzo Mikami, Wataru Hashimoto, Kousaku Murata
Structural determinants responsible for substrate recognition and mode of action in family 11 polysaccharide lyases.
J Biol Chem: 2009, 284(15);10181-9
[PubMed:19193638]
[WorldCat.org]
[DOI]
(P p)
Akihito Ochiai, Takafumi Itoh, Yukie Maruyama, Akiko Kawamata, Bunzo Mikami, Wataru Hashimoto, Kousaku Murata
A novel structural fold in polysaccharide lyases: Bacillus subtilis family 11 rhamnogalacturonan lyase YesW with an eight-bladed beta-propeller.
J Biol Chem: 2007, 282(51);37134-45
[PubMed:17947240]
[WorldCat.org]
[DOI]
(P p)
Akihito Ochiai, Takafumi Itoh, Akiko Kawamata, Wataru Hashimoto, Kousaku Murata
Plant cell wall degradation by saprophytic Bacillus subtilis strains: gene clusters responsible for rhamnogalacturonan depolymerization.
Appl Environ Microbiol: 2007, 73(12);3803-13
[PubMed:17449691]
[WorldCat.org]
[DOI]
(P p)
Akihito Ochiai, Masayuki Yamasaki, Takafumi Itoh, Bunzo Mikami, Wataru Hashimoto, Kousaku Murata
Crystallization and preliminary X-ray analysis of the rhamnogalacturonan lyase YesW from Bacillus subtilis strain 168, a member of polysaccharide lyase family 11.
Acta Crystallogr Sect F Struct Biol Cryst Commun: 2006, 62(Pt 5);438-40
[PubMed:16682770]
[WorldCat.org]
[DOI]
(I p)