Difference between revisions of "YcgT"
(→Additional information) |
|||
| Line 86: | Line 86: | ||
=== Additional information=== | === Additional information=== | ||
The gene is annotated in KEGG as an ortholog of thioredoxin reductase (NADPH) EC 1.8.1.9. In MetaCyc the protein is marked as “similar to thioredoxin | The gene is annotated in KEGG as an ortholog of thioredoxin reductase (NADPH) EC 1.8.1.9. In MetaCyc the protein is marked as “similar to thioredoxin | ||
| − | reductase”. No EC annotation is available in Swiss-Prot. No literature/experimental evidence supporting the annotation is | + | reductase”. No EC annotation is available in Swiss-Prot. No literature/experimental evidence supporting the annotation is available. {{PubMed|19935659}} |
=Expression and regulation= | =Expression and regulation= | ||
Revision as of 11:48, 26 November 2009
- Description: unknown
| Gene name | ycgT |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | unknown |
| MW, pI | 36 kDa, 5.61 |
| Gene length, protein length | 1008 bp, 336 aa |
| Immediate neighbours | ycgS, nasF |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 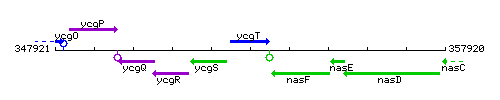
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU03270
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 2 reduced ferredoxin + NADP+ + H+ = 2 oxidized ferredoxin + NADPH (according to Swiss-Prot)
- Protein family: ferredoxin--NADP reductase type 2 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: O31475
- KEGG entry: [3]
- E.C. number:
Additional information
The gene is annotated in KEGG as an ortholog of thioredoxin reductase (NADPH) EC 1.8.1.9. In MetaCyc the protein is marked as “similar to thioredoxin reductase”. No EC annotation is available in Swiss-Prot. No literature/experimental evidence supporting the annotation is available. PubMed
Expression and regulation
- Operon: ycgT (according to DBTBS)
- Additional information: the mRNA is very stable (half-life > 15 min) PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Juliane Ollinger, Kyung-Bok Song, Haike Antelmann, Michael Hecker, John D Helmann
Role of the Fur regulon in iron transport in Bacillus subtilis.
J Bacteriol: 2006, 188(10);3664-73
[PubMed:16672620]
[WorldCat.org]
[DOI]
(P p)
G Hambraeus, C von Wachenfeldt, L Hederstedt
Genome-wide survey of mRNA half-lives in Bacillus subtilis identifies extremely stable mRNAs.
Mol Genet Genomics: 2003, 269(5);706-14
[PubMed:12884008]
[WorldCat.org]
[DOI]
(P p)