Difference between revisions of "YacL"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 100: | Line 100: | ||
* '''Operon:''' ''[[yacL]]-[[ispD]]-[[ispF]]'' {{PubMed|10482513}} | * '''Operon:''' ''[[yacL]]-[[ispD]]-[[ispF]]'' {{PubMed|10482513}} | ||
| − | * '''[ | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=yacL_108674_109774_1 yacL] {{PubMed|22383849}} |
| + | |||
| + | * '''Sigma factor:''' [[SigB]] [http://www.ncbi.nlm.nih.gov/pubmed/15805528 PubMed], [[SigM]] [http://www.ncbi.nlm.nih.gov/sites/entrez/17434969 PubMed] | ||
* '''Regulation:''' induced by stress ([[SigB]]) [http://www.ncbi.nlm.nih.gov/pubmed/15805528 PubMed] | * '''Regulation:''' induced by stress ([[SigB]]) [http://www.ncbi.nlm.nih.gov/pubmed/15805528 PubMed] | ||
Revision as of 14:35, 7 August 2012
- Description: general stress protein, survival of salt and ethanol stresses
| Gene name | yacL |
| Synonyms | |
| Essential | no |
| Product | unknown |
| Function | survival of salt and ethanol stresses |
| MW, pI | 40 kDa, 4.97 |
| Gene length, protein length | 1098 bp, 366 aa |
| Immediate neighbours | disA, ispD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 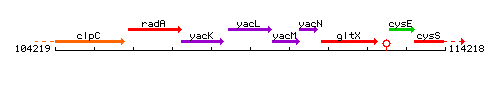
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
general stress proteins (controlled by SigB), cell envelope stress proteins (controlled by SigM, V, W, X, Y)
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU00890
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: TRAM domain (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: Q06754
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Adrian J Jervis, Penny D Thackray, Chris W Houston, Malcolm J Horsburgh, Anne Moir
SigM-responsive genes of Bacillus subtilis and their promoters.
J Bacteriol: 2007, 189(12);4534-8
[PubMed:17434969]
[WorldCat.org]
[DOI]
(P p)
Dirk Höper, Uwe Völker, Michael Hecker
Comprehensive characterization of the contribution of individual SigB-dependent general stress genes to stress resistance of Bacillus subtilis.
J Bacteriol: 2005, 187(8);2810-26
[PubMed:15805528]
[WorldCat.org]
[DOI]
(P p)
A Petersohn, J Bernhardt, U Gerth, D Höper, T Koburger, U Völker, M Hecker
Identification of sigma(B)-dependent genes in Bacillus subtilis using a promoter consensus-directed search and oligonucleotide hybridization.
J Bacteriol: 1999, 181(18);5718-24
[PubMed:10482513]
[WorldCat.org]
[DOI]
(P p)