Difference between revisions of "TufA"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 47: | Line 47: | ||
* '''Locus tag:''' BSU01130 | * '''Locus tag:''' BSU01130 | ||
| − | |||
| − | |||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
Revision as of 17:28, 27 January 2012
- Description: elongation factor Tu
| Gene name | tufA |
| Synonyms | |
| Essential | yes PubMed |
| Product | elongation factor Tu |
| Function | translation |
| Interactions involving this protein in SubtInteract: TufA | |
| MW, pI | 43 kDa, 4.723 |
| Gene length, protein length | 1188 bp, 396 aa |
| Immediate neighbours | fusA, ybaC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 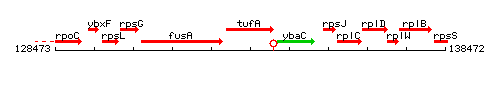
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
translation, essential genes, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU01130
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: EF-Tu/EF-1A subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P33166
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Hervé Joël Defeu Soufo, Christian Reimold, Uwe Linne, Tobias Knust, Johannes Gescher, Peter L Graumann
Bacterial translation elongation factor EF-Tu interacts and colocalizes with actin-like MreB protein.
Proc Natl Acad Sci U S A: 2010, 107(7);3163-8
[PubMed:20133608]
[WorldCat.org]
[DOI]
(I p)
Cédric Absalon, Michal Obuchowski, Edwige Madec, Delphine Delattre, I Barry Holland, Simone J Séror
CpgA, EF-Tu and the stressosome protein YezB are substrates of the Ser/Thr kinase/phosphatase couple, PrkC/PrpC, in Bacillus subtilis.
Microbiology (Reading): 2009, 155(Pt 3);932-943
[PubMed:19246764]
[WorldCat.org]
[DOI]
(P p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Christine Eymann, Dörte Becher, Jörg Bernhardt, Katrin Gronau, Anja Klutzny, Michael Hecker
Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis.
Proteomics: 2007, 7(19);3509-26
[PubMed:17726680]
[WorldCat.org]
[DOI]
(P p)