Tig
- Description: trigger factor (prolyl isomerase)
| Gene name | tig |
| Synonyms | yzzH |
| Essential | no |
| Product | trigger factor (prolyl isomerase) |
| Function | protein folding |
| Gene expression levels in SubtiExpress: tig | |
| Interactions involving this protein in SubtInteract: Tig | |
| MW, pI | 47 kDa, 4.224 |
| Gene length, protein length | 1272 bp, 424 aa |
| Immediate neighbours | clpX, ysoA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 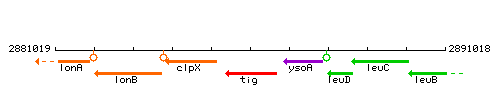
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed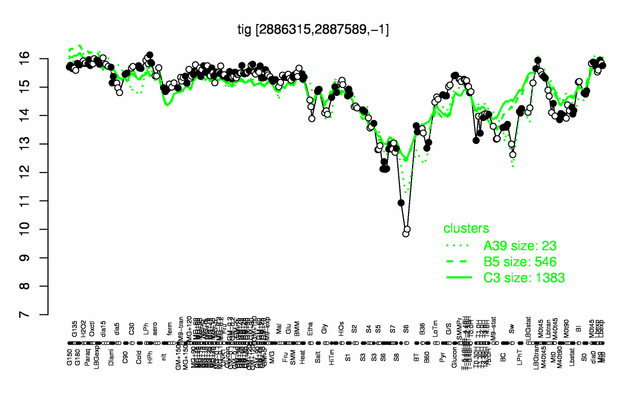
| |
Contents
Categories containing this gene/protein
chaperones/ protein folding, phosphoproteins, most abundant proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28230
Phenotypes of a mutant
Database entries
- BsubCyc: BSU28230
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: Tig subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Effectors of protein activity:
Database entries
- BsubCyc: BSU28230
- UniProt: P80698
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon: tig PubMed
- Regulatory mechanism:
- Additional information:
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
- The mRNA has a long 5' leader region. This may indicate RNA-based regulation PubMed
- belongs to the 100 most abundant proteins PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium): 9318 PubMed
- number of protein molecules per cell (complex medium with amino acids, without glucose): 56291 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Irnov Irnov, Cynthia M Sharma, Jörg Vogel, Wade C Winkler
Identification of regulatory RNAs in Bacillus subtilis.
Nucleic Acids Res: 2010, 38(19);6637-51
[PubMed:20525796]
[WorldCat.org]
[DOI]
(I p)
Frieder Merz, Daniel Boehringer, Christiane Schaffitzel, Steffen Preissler, Anja Hoffmann, Timm Maier, Anna Rutkowska, Jasmin Lozza, Nenad Ban, Bernd Bukau, Elke Deuerling
Molecular mechanism and structure of Trigger Factor bound to the translating ribosome.
EMBO J: 2008, 27(11);1622-32
[PubMed:18497744]
[WorldCat.org]
[DOI]
(I p)
Alain Lévine, Françoise Vannier, Cédric Absalon, Lauriane Kuhn, Peter Jackson, Elaine Scrivener, Valérie Labas, Joëlle Vinh, Patrick Courtney, Jérôme Garin, Simone J Séror
Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes.
Proteomics: 2006, 6(7);2157-73
[PubMed:16493705]
[WorldCat.org]
[DOI]
(P p)
Frank Schlünzen, Daniel N Wilson, Pingsheng Tian, Jörg M Harms, Stuart J McInnes, Harly A S Hansen, Renate Albrecht, Jörg Buerger, Sigurd M Wilbanks, Paola Fucini
The binding mode of the trigger factor on the ribosome: implications for protein folding and SRP interaction.
Structure: 2005, 13(11);1685-94
[PubMed:16271892]
[WorldCat.org]
[DOI]
(P p)
David Baram, Erez Pyetan, Assa Sittner, Tamar Auerbach-Nevo, Anat Bashan, Ada Yonath
Structure of trigger factor binding domain in biologically homologous complex with eubacterial ribosome reveals its chaperone action.
Proc Natl Acad Sci U S A: 2005, 102(34);12017-22
[PubMed:16091460]
[WorldCat.org]
[DOI]
(P p)
Christine Eymann, Annette Dreisbach, Dirk Albrecht, Jörg Bernhardt, Dörte Becher, Sandy Gentner, Le Thi Tam, Knut Büttner, Gerrit Buurman, Christian Scharf, Simone Venz, Uwe Völker, Michael Hecker
A comprehensive proteome map of growing Bacillus subtilis cells.
Proteomics: 2004, 4(10);2849-76
[PubMed:15378759]
[WorldCat.org]
[DOI]
(P p)
Günter Kramer, Thomas Rauch, Wolfgang Rist, Sonja Vorderwülbecke, Holger Patzelt, Agnes Schulze-Specking, Nenad Ban, Elke Deuerling, Bernd Bukau
L23 protein functions as a chaperone docking site on the ribosome.
Nature: 2002, 419(6903);171-4
[PubMed:12226666]
[WorldCat.org]
[DOI]
(P p)
Dindo Y Reyes, Hirofumi Yoshikawa
DnaK chaperone machine and trigger factor are only partially required for normal growth of Bacillus subtilis.
Biosci Biotechnol Biochem: 2002, 66(7);1583-6
[PubMed:12224648]
[WorldCat.org]
[DOI]
(P p)
S F Göthel, C Scholz, F X Schmid, M A Marahiel
Cyclophilin and trigger factor from Bacillus subtilis catalyze in vitro protein folding and are necessary for viability under starvation conditions.
Biochemistry: 1998, 37(38);13392-9
[PubMed:9748346]
[WorldCat.org]
[DOI]
(P p)
S F Göthel, R Schmid, A Wipat, N M Carter, P T Emmerson, C R Harwood, M A Marahiel
An internal FK506-binding domain is the catalytic core of the prolyl isomerase activity associated with the Bacillus subtilis trigger factor.
Eur J Biochem: 1997, 244(1);59-65
[PubMed:9063446]
[WorldCat.org]
[DOI]
(P p)
A Wipat, N Carter, S C Brignell, B J Guy, K Piper, J Sanders, P T Emmerson, C R Harwood
The dnaB-pheA (256 degrees-240 degrees) region of the Bacillus subtilis chromosome containing genes responsible for stress responses, the utilization of plant cell walls and primary metabolism.
Microbiology (Reading): 1996, 142 ( Pt 11);3067-78
[PubMed:8969504]
[WorldCat.org]
[DOI]
(P p)