Difference between revisions of "TagE"
(→Extended information on the protein) |
|||
| Line 99: | Line 99: | ||
* '''Operon:''' ''[[tagD]]-[[tagE]]-[[tagF]]'' {{PubMed|2507871}} | * '''Operon:''' ''[[tagD]]-[[tagE]]-[[tagF]]'' {{PubMed|2507871}} | ||
| − | * '''[ | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=tagE_3678399_3680420_-1 tagE] {{PubMed|22383849}} |
| + | |||
| + | * '''Sigma factor:''' [[SigA]] {{PubMed|7581998}} | ||
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 108: | Line 110: | ||
** [[WalR]]: transcription activation {{PubMed|12950927}} | ** [[WalR]]: transcription activation {{PubMed|12950927}} | ||
| − | * '''Additional information:''' | + | * '''Additional information:''' |
=Biological materials = | =Biological materials = | ||
Revision as of 07:46, 17 April 2012
- Description: UDP-glucose:polyglycerol phosphate glucosyltransferase
| Gene name | tagE |
| Synonyms | rodD, gtaA, gtaD |
| Essential | no |
| Product | UDP-glucose:polyglycerol phosphate glucosyltransferase |
| Function | biosynthesis of teichoic acid |
| MW, pI | 78 kDa, 6.026 |
| Gene length, protein length | 2019 bp, 673 aa |
| Immediate neighbours | tagF, tagD |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 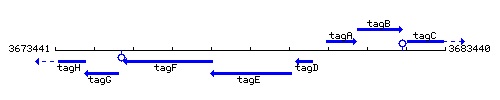
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
cell wall synthesis, biosynthesis of cell wall components, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU35730
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transfer of glucose from UDP-glucose to glycerol phosphate in polymeric form PubMed
- Protein family: glycosyltransferase 1 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation on Ser-2 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: P13484
- KEGG entry: [3]
- E.C. number: 2.4.1.52
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Boris Macek, Ivan Mijakovic, Jesper V Olsen, Florian Gnad, Chanchal Kumar, Peter R Jensen, Matthias Mann
The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis.
Mol Cell Proteomics: 2007, 6(4);697-707
[PubMed:17218307]
[WorldCat.org]
[DOI]
(P p)
Alistair Howell, Sarah Dubrac, Kasper Krogh Andersen, David Noone, Juliette Fert, Tarek Msadek, Kevin Devine
Genes controlled by the essential YycG/YycF two-component system of Bacillus subtilis revealed through a novel hybrid regulator approach.
Mol Microbiol: 2003, 49(6);1639-55
[PubMed:12950927]
[WorldCat.org]
[DOI]
(P p)
W Liu, S Eder, F M Hulett
Analysis of Bacillus subtilis tagAB and tagDEF expression during phosphate starvation identifies a repressor role for PhoP-P.
J Bacteriol: 1998, 180(3);753-8
[PubMed:9457886]
[WorldCat.org]
[DOI]
(P p)
C Mauël, A Bauduret, C Chervet, S Beggah, D Karamata
In Bacillus subtilis 168, teichoic acid of the cross-wall may be different from that of the cylinder: a hypothesis based on transcription analysis of tag genes.
Microbiology (Reading): 1995, 141 ( Pt 10);2379-89
[PubMed:7581998]
[WorldCat.org]
[DOI]
(P p)
A L Honeyman, G C Stewart
The nucleotide sequence of the rodC operon of Bacillus subtilis.
Mol Microbiol: 1989, 3(9);1257-68
[PubMed:2507871]
[WorldCat.org]
[DOI]
(P p)