SndA
- Description: sulphur compound N-deacetylase, required for the conversion of S-methyl-cysteine to cysteine
| Gene name | sndA |
| Synonyms | hipO, ytnL |
| Essential | no |
| Product | sulphur compound N-deacetylase |
| Function | utilization of S-methyl-cysteine |
| Gene expression levels in SubtiExpress: sndA | |
| Metabolic function and regulation of this protein in SubtiPathways: Riboflavin / FAD | |
| MW, pI | 45 kDa, 5.545 |
| Gene length, protein length | 1248 bp, 416 aa |
| Immediate neighbours | ytnM, ribR |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 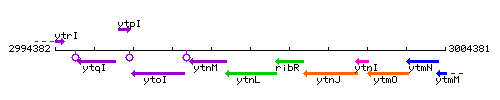
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed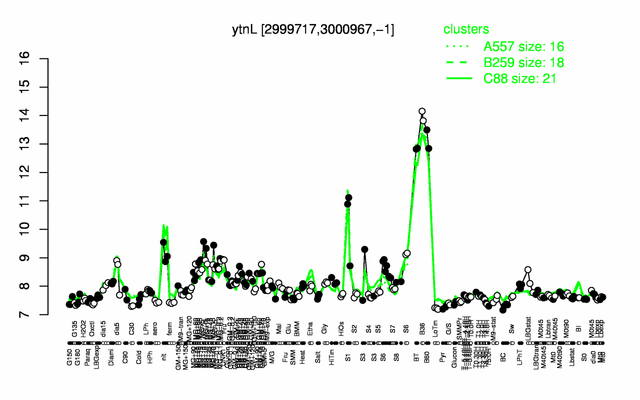
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
CymR regulon, AscR regulon, Efp-dependent proteins
The gene
Basic information
- Locus tag: BSU29290
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: peptidase M20 family (according to Swiss-Prot)
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O34980
- KEGG entry: [3]
- E.C. number:
Additional information
The gene was annotated in KEGG and MetaCyc as hippurate hydrolase EC 3.5.1.32. In Swiss-Prot it is marked as “uncharacterized hydrolase”. No literature/experimental evidence supporting ththeannotation was removed as of November 2009. PubMed
Expression and regulation
- Regulation:
- Additional information:
- translation is likely to require Efp due to the presence of several consecutive proline residues PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Che-Man Chan, Antoine Danchin, Philippe Marlière, Agnieszka Sekowska
Paralogous metabolism: S-alkyl-cysteine degradation in Bacillus subtilis.
Environ Microbiol: 2014, 16(1);101-17
[PubMed:23944997]
[WorldCat.org]
[DOI]
(I p)
Pierre Burguière, Juliette Fert, Isabelle Guillouard, Sandrine Auger, Antoine Danchin, Isabelle Martin-Verstraete
Regulation of the Bacillus subtilis ytmI operon, involved in sulfur metabolism.
J Bacteriol: 2005, 187(17);6019-30
[PubMed:16109943]
[WorldCat.org]
[DOI]
(P p)
Irina M Solovieva, Rimma A Kreneva, Lubov Errais Lopes, Daniel A Perumov
The riboflavin kinase encoding gene ribR of Bacillus subtilis is a part of a 10 kb operon, which is negatively regulated by the yrzC gene product.
FEMS Microbiol Lett: 2005, 243(1);51-8
[PubMed:15668000]
[WorldCat.org]
[DOI]
(P p)
I M Solov'eva, R A Kreneva, L L Erraĭs, A S Mironov, D A Perumov
[Study of the mechanism for regulating ribR gene activity in Bacillus subtilis]. [Issledovanie mekhanizma reguliatsii aktivnosti gena ribR u Bacillus subtilis.]
Genetika: 2004, 40(5);716-20
[PubMed:15272571]
[WorldCat.org]
(P p)