Difference between revisions of "SlrR"
| Line 159: | Line 159: | ||
<pubmed> 20541494 23353768 </pubmed> | <pubmed> 20541494 23353768 </pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | + | <pubmed>23378512 23430750 21856853 22329926,22113911,20817675 18430133,18647168, 19767430, 19788541 20351052 20923420 </pubmed> | |
| − | <pubmed>23378512 23430750 | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:49, 27 March 2013
- Description: transcriptional activator of competence development and sporulation genes, represses SigD-dependent flagellar genes, antagonist of SlrA and SinR, has LexA-like autocleavage activity
| Gene name | slrR |
| Synonyms | yveJ, slr |
| Essential | no |
| Product | transcription regulator, SlrA antagonist |
| Function | regulation of initiation of biofilm formation and of autolysis |
| Gene expression levels in SubtiExpress: slrR | |
| Interactions involving this protein in SubtInteract: SlrR | |
| Regulation of this protein in SubtiPathways: Biofilm | |
| MW, pI | 17 kDa, 9.63 |
| Gene length, protein length | 456 bp, 152 aa |
| Immediate neighbours | epsA, pnbA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 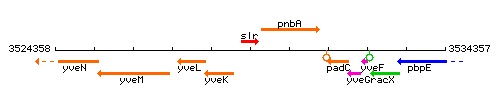
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed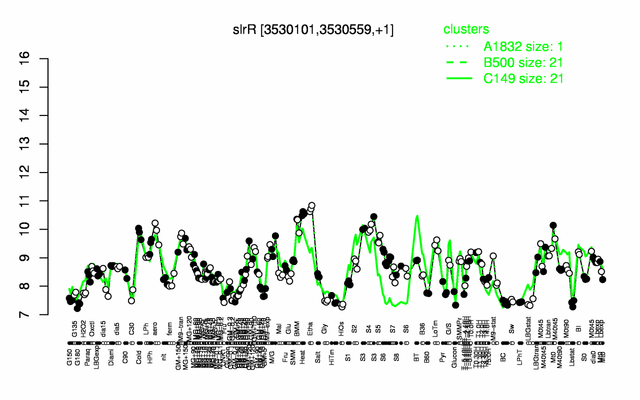
| |
Contents
Categories containing this gene/protein
transcription factors and their control, transition state regulators, biofilm formation
This gene is a member of the following regulons
Abh regulon, AbrB regulon, SinR regulon
The SlrR regulon:
The gene
Basic information
- Locus tag: BSU34380
Phenotypes of a mutant
- smooth colonies on MsGG medium, no biofilm formation PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- SlrR binds to and inhibits the activity of SlrA, SlrA indirectly stimulates the synthesis of SlrR by interacting with SinR. SlrR can bind to SinR and SinR directly represses the transcription of SlrR. SlrR indirectly derepresses its own gene. The heterocomplex of SlrR-SinR is a repressor of autolysin and motility genes and inhibits the repressor function of SinR. PubMed
- repression of transcription of lytA-lytB-lytC and lytF PubMed
- autocleavage PubMed
- Protein family:
- Paralogous protein(s): SinR
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity: interaction with SinR triggers binding of SlrR to the promoters of lytA-lytB-lytC and lytF, resulting in their repression PubMed
Database entries
- Structure:
- UniProt: P71049
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- GP955 (slrR-pnbA::cat), available in Jörg Stülke's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Hera Vlamakis, Yunrong Chai, Pascale Beauregard, Richard Losick, Roberto Kolter
Sticking together: building a biofilm the Bacillus subtilis way.
Nat Rev Microbiol: 2013, 11(3);157-68
[PubMed:23353768]
[WorldCat.org]
[DOI]
(I p)
Patrick Piggot
Epigenetic switching: bacteria hedge bets about staying or moving.
Curr Biol: 2010, 20(11);R480-2
[PubMed:20541494]
[WorldCat.org]
[DOI]
(I p)
Original publications
Joseph A Newman, Cecilia Rodrigues, Richard J Lewis
Molecular basis of the activity of SinR protein, the master regulator of biofilm formation in Bacillus subtilis.
J Biol Chem: 2013, 288(15);10766-78
[PubMed:23430750]
[WorldCat.org]
[DOI]
(I p)
Ying Lei, Taku Oshima, Naotake Ogasawara, Shu Ishikawa
Functional analysis of the protein Veg, which stimulates biofilm formation in Bacillus subtilis.
J Bacteriol: 2013, 195(8);1697-705
[PubMed:23378512]
[WorldCat.org]
[DOI]
(I p)
Loralyn M Cozy, Andrew M Phillips, Rebecca A Calvo, Ashley R Bate, Yi-Huang Hsueh, Richard Bonneau, Patrick Eichenberger, Daniel B Kearns
SlrA/SinR/SlrR inhibits motility gene expression upstream of a hypersensitive and hysteretic switch at the level of σ(D) in Bacillus subtilis.
Mol Microbiol: 2012, 83(6);1210-28
[PubMed:22329926]
[WorldCat.org]
[DOI]
(I p)
Eric R Pozsgai, Kris M Blair, Daniel B Kearns
Modified mariner transposons for random inducible-expression insertions and transcriptional reporter fusion insertions in Bacillus subtilis.
Appl Environ Microbiol: 2012, 78(3);778-85
[PubMed:22113911]
[WorldCat.org]
[DOI]
(I p)
Christine Diethmaier, Nico Pietack, Katrin Gunka, Christoph Wrede, Martin Lehnik-Habrink, Christina Herzberg, Sebastian Hübner, Jörg Stülke
A novel factor controlling bistability in Bacillus subtilis: the YmdB protein affects flagellin expression and biofilm formation.
J Bacteriol: 2011, 193(21);5997-6007
[PubMed:21856853]
[WorldCat.org]
[DOI]
(I p)
Yunrong Chai, Roberto Kolter, Richard Losick
Reversal of an epigenetic switch governing cell chaining in Bacillus subtilis by protein instability.
Mol Microbiol: 2010, 78(1);218-29
[PubMed:20923420]
[WorldCat.org]
[DOI]
(I p)
Onuma Chumsakul, Hiroki Takahashi, Taku Oshima, Takahiro Hishimoto, Shigehiko Kanaya, Naotake Ogasawara, Shu Ishikawa
Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation.
Nucleic Acids Res: 2011, 39(2);414-28
[PubMed:20817675]
[WorldCat.org]
[DOI]
(I p)
Yunrong Chai, Thomas Norman, Roberto Kolter, Richard Losick
An epigenetic switch governing daughter cell separation in Bacillus subtilis.
Genes Dev: 2010, 24(8);754-65
[PubMed:20351052]
[WorldCat.org]
[DOI]
(I p)
Yunrong Chai, Roberto Kolter, Richard Losick
Paralogous antirepressors acting on the master regulator for biofilm formation in Bacillus subtilis.
Mol Microbiol: 2009, 74(4);876-87
[PubMed:19788541]
[WorldCat.org]
[DOI]
(I p)
Ewan J Murray, Mark A Strauch, Nicola R Stanley-Wall
SigmaX is involved in controlling Bacillus subtilis biofilm architecture through the AbrB homologue Abh.
J Bacteriol: 2009, 191(22);6822-32
[PubMed:19767430]
[WorldCat.org]
[DOI]
(I p)
Kazuo Kobayashi
SlrR/SlrA controls the initiation of biofilm formation in Bacillus subtilis.
Mol Microbiol: 2008, 69(6);1399-410
[PubMed:18647168]
[WorldCat.org]
[DOI]
(I p)
Frances Chu, Daniel B Kearns, Anna McLoon, Yunrong Chai, Roberto Kolter, Richard Losick
A novel regulatory protein governing biofilm formation in Bacillus subtilis.
Mol Microbiol: 2008, 68(5);1117-27
[PubMed:18430133]
[WorldCat.org]
[DOI]
(I p)