Difference between revisions of "Sandbox"
Raphael2215 (talk | contribs) |
Raphael2215 (talk | contribs) |
||
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' D-alanine transfer from Dcp to undecaprenol-phosphate <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''dltB'' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' '' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || ''ipa-4r '' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | + | |style="background:#ABCDEF;" align="center"| '''Essential''' || no |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || D-alanine transfer from Dcp to undecaprenol-phosphate |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || biosynthesis of teichoic acid |
| + | acid (D-alanyl transfer from Dcp to undecaprenol-phosphate) | ||
|- | |- | ||
| − | | | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 46 kDa, 9.944 |
|- | |- | ||
| − | | | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1185 bp, 395 aa |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| ''' | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[dltA]]'', ''[[dltC]]'' |
|- | |- | ||
| − | |style="background:# | + | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB15877]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' |
|- | |- | ||
| − | + | |colspan="2" | '''Genetic context''' <br/> [[Image:dltB_context.gif]] | |
| − | |||
| − | |||
| − | |||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
|- | |- | ||
| − | |colspan="2" |'''Expression'''<br/>[[Image: | + | |colspan="2" |'''Expression'''<br/>[[Image:dltB_expression.png|400px]]<div align="right"> <small>[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=dltB_3953783_3954970_1 Expression at a glance]</small></div> |
|} | |} | ||
| − | |||
| − | |||
| − | |||
| − | |||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| Line 40: | Line 33: | ||
__TOC__ | __TOC__ | ||
| − | + | <br/><br/> | |
| − | |||
| − | <br/><br/> | ||
| − | |||
= [[Categories]] containing this gene/protein = | = [[Categories]] containing this gene/protein = | ||
| − | {{SubtiWiki category|[[ | + | {{SubtiWiki category|[[cell wall synthesis]]}}, |
| − | {{SubtiWiki category|[[ | + | {{SubtiWiki category|[[transporters/ other]]}}, |
| − | {{SubtiWiki category|[[ | + | {{SubtiWiki category|[[biosynthesis of cell wall components]]}}, |
| − | {{SubtiWiki category|[[ | + | {{SubtiWiki category|[[cell envelope stress proteins (controlled by SigM, V, W, X, Y)]]}}, |
| + | {{SubtiWiki category|[[membrane proteins]]}} | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
| − | {{SubtiWiki regulon|[[ | + | {{SubtiWiki regulon|[[SigD regulon]]}}, |
| + | {{SubtiWiki regulon|[[SigM regulon]]}}, | ||
| + | {{SubtiWiki regulon|[[SigX regulon]]}}, | ||
| + | {{SubtiWiki regulon|[[Spo0A regulon]]}}, | ||
| + | {{SubtiWiki regulon|[[stringent response]]}}, | ||
| + | {{SubtiWiki regulon|[[YvrHb regulon]]}} | ||
=The gene= | =The gene= | ||
=== Basic information === | === Basic information === | ||
| − | * '''Locus tag:''' | + | |
| + | * '''Locus tag:''' BSU38510 | ||
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | |||
| − | |||
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ | + | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/dltABCDE.html] |
| − | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG10550] |
=== Additional information=== | === Additional information=== | ||
| + | |||
| + | |||
| + | |||
=The protein= | =The protein= | ||
| Line 75: | Line 73: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' |
| − | * '''Protein family:''' | + | * '''Protein family:''' membrane-bound acyltransferase family (according to Swiss-Prot) |
* '''Paralogous protein(s):''' | * '''Paralogous protein(s):''' | ||
| Line 83: | Line 81: | ||
=== Extended information on the protein === | === Extended information on the protein === | ||
| − | * '''Kinetic information:''' | + | * '''Kinetic information:''' |
* '''Domains:''' | * '''Domains:''' | ||
| − | |||
| − | * '''Modification:''' | + | * '''Modification:''' |
| − | * '''Cofactor(s):''' | + | * '''Cofactor(s):''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | |||
* '''[[SubtInteract|Interactions]]:''' | * '''[[SubtInteract|Interactions]]:''' | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | * '''[[Localization]]:''' | + | * '''[[Localization]]:''' |
| − | + | ** membrane associated [http://www.ncbi.nlm.nih.gov/pubmed/18763711 PubMed] | |
| − | ** membrane associated [http://www.ncbi.nlm.nih.gov/ | ||
| − | |||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' |
| − | |||
| − | |||
| − | * '''UniProt:''' [http://www.uniprot.org/uniprot/ | + | * '''UniProt:''' [http://www.uniprot.org/uniprot/P39580 P39580] |
| − | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu: | + | * '''KEGG entry:''' [http://www.genome.jp/dbget-bin/www_bget?bsu:BSU38510] |
| − | * '''E.C. number:''' | + | * '''E.C. number:''' |
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
| − | |||
=Expression and regulation= | =Expression and regulation= | ||
| + | * '''Operon:''' ''[[dltA]]-[[dltB]]-[[dltC]]-[[dltD]]-[[dltE]]'' {{PubMed|7797557}} | ||
| − | * ''' | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=dltB_3953783_3954970_1 dltB] {{PubMed|22383849}} |
| − | |||
| − | |||
| − | * ''' | + | * '''Sigma factor:''' [[SigD]] {{PubMed|7797557}}, [[SigM]] {{PubMed|18179421}}, [[SigX]] {{PubMed|14762009}} |
| − | |||
| − | |||
* '''Regulation:''' | * '''Regulation:''' | ||
| − | ** | + | ** [[RelA]] dependent downregulation (Class I) during stringent response {{PubMed|11948165}} |
| − | ** | + | ** repressed under conditions that trigger sporulation ([[Spo0A]]) {{PubMed|7797557}} |
| − | ** | + | ** expression is reduced in a [[SigV]] mutant {{PubMed|21926231}} |
| − | * '''Regulatory mechanism:''' | + | * '''Regulatory mechanism:''' |
| + | ** activated by [[YvrHb]] [http://www.ncbi.nlm.nih.gov/sites/entrez/16306698 PubMed] | ||
| + | ** [[Spo0A]]: transcription repression {{PubMed|7797557}} | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| Line 146: | Line 128: | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' | + | * '''Mutant:''' |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| + | * '''Expression vector:''' | ||
| + | |||
* '''lacZ fusion:''' | * '''lacZ fusion:''' | ||
| − | |||
| − | |||
| − | |||
| − | * ''' | + | * '''GFP fusion:''' |
| − | * ''' | + | * '''two-hybrid system:''' |
| − | * '''Antibody:''' | + | * '''Antibody:''' |
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
| − | |||
| − | |||
| − | |||
=Your additional remarks= | =Your additional remarks= | ||
=References= | =References= | ||
| − | + | '''Additional publications:''' {{PubMed|21856855,21926231}} | |
| − | + | <pubmed>14762009,12028417 ,7797557,18763711, </pubmed> | |
| − | |||
| − | '''Additional publications:''' {{PubMed| | ||
| − | <pubmed> | ||
| − | |||
| − | |||
| − | |||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:46, 18 April 2012
- Description: D-alanine transfer from Dcp to undecaprenol-phosphate
| Gene name | dltB |
| Synonyms | ipa-4r |
| Essential | no |
| Product | D-alanine transfer from Dcp to undecaprenol-phosphate |
| Function | biosynthesis of teichoic acid
acid (D-alanyl transfer from Dcp to undecaprenol-phosphate) |
| MW, pI | 46 kDa, 9.944 |
| Gene length, protein length | 1185 bp, 395 aa |
| Immediate neighbours | dltA, dltC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 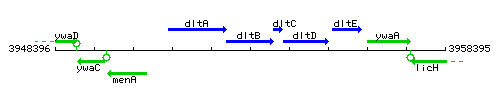
This image was kindly provided by SubtiList
| |
Expression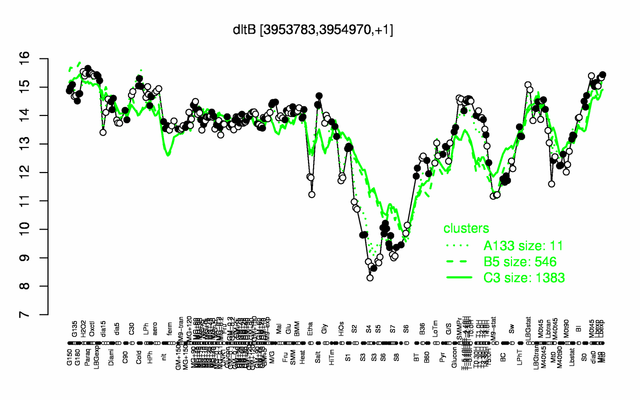
| |
Contents
Categories containing this gene/protein
cell wall synthesis, transporters/ other, biosynthesis of cell wall components, cell envelope stress proteins (controlled by SigM, V, W, X, Y), membrane proteins
This gene is a member of the following regulons
SigD regulon, SigM regulon, SigX regulon, Spo0A regulon, stringent response, YvrHb regulon
The gene
Basic information
- Locus tag: BSU38510
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: membrane-bound acyltransferase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- membrane associated PubMed
Database entries
- Structure:
- UniProt: P39580
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Min Cao, John D Helmann
The Bacillus subtilis extracytoplasmic-function sigmaX factor regulates modification of the cell envelope and resistance to cationic antimicrobial peptides.
J Bacteriol: 2004, 186(4);1136-46
[PubMed:14762009]
[WorldCat.org]
[DOI]
(P p)
K Stephenson, C L Jensen, S T Jørgensen, C R Harwood
Simultaneous inactivation of the wprA and dltB genes of Bacillus subtilis reduces the yield of alpha-amylase.
Lett Appl Microbiol: 2002, 34(6);394-7
[PubMed:12028417]
[WorldCat.org]
[DOI]
(P p)
M Perego, P Glaser, A Minutello, M A Strauch, K Leopold, W Fischer
Incorporation of D-alanine into lipoteichoic acid and wall teichoic acid in Bacillus subtilis. Identification of genes and regulation.
J Biol Chem: 1995, 270(26);15598-606
[PubMed:7797557]
[WorldCat.org]
[DOI]
(P p)