Difference between revisions of "Sandbox"
| Line 1: | Line 1: | ||
| − | * '''Description:''' | + | * '''Description:''' replication initiation protein <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Gene name''' | |style="background:#ABCDEF;" align="center"|'''Gene name''' | ||
| − | |'' | + | |''dnaA'' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || '' '' | + | |style="background:#ABCDEF;" align="center"| '''Synonyms''' || ''dnaH, dnaJ, dnaK '' |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Essential''' || | + | |style="background:#ABCDEF;" align="center"| '''Essential''' || yes |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || | + | |style="background:#ABCDEF;" align="center"| '''Product''' || |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || | + | |style="background:#ABCDEF;" align="center"|'''Function''' || initiation of chromosome replication |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || | + | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 50 kDa, 6.035 |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || | + | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1338 bp, 446 aa |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || |
|- | |- | ||
| − | |style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/ | + | |style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/dnaA_nucleotide.txt Gene sequence (+200bp) ]''' |
| − | |style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/ | + | |style="background:#FAF8CC;" align="center"|'''[http://subtiwiki.uni-goettingen.de/dnaA_protein.txt Protein sequence]''' |
|- | |- | ||
| − | |colspan="2" | '''Genetic context''' <br/> [[Image: | + | |colspan="2" | '''Genetic context''' <br/> [[Image:dnaA_context.gif]] |
|- | |- | ||
|} | |} | ||
| Line 31: | Line 31: | ||
<br/><br/> | <br/><br/> | ||
| − | |||
=The gene= | =The gene= | ||
=== Basic information === | === Basic information === | ||
| − | * '''Coordinates:''' | + | * '''Coordinates:''' |
===Phenotypes of a mutant === | ===Phenotypes of a mutant === | ||
| − | + | essential | |
| − | essential | ||
=== Database entries === | === Database entries === | ||
| − | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ | + | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/dnaAN.html] |
| − | * '''SubtiList entry:'''[http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+ | + | * '''SubtiList entry:''' [http://genolist.pasteur.fr/SubtiList/genome.cgi?gene_detail+BG10065] |
=== Additional information=== | === Additional information=== | ||
| + | |||
=The protein= | =The protein= | ||
| Line 54: | Line 53: | ||
=== Basic information/ Evolution === | === Basic information/ Evolution === | ||
| − | * '''Catalyzed reaction/ biological activity:''' | + | * '''Catalyzed reaction/ biological activity:''' |
* '''Protein family:''' | * '''Protein family:''' | ||
| − | * '''Paralogous protein(s):''' | + | * '''Paralogous protein(s):''' |
=== Extended information on the protein === | === Extended information on the protein === | ||
| − | * '''Kinetic information:''' | + | * '''Kinetic information:''' |
* '''Domains:''' | * '''Domains:''' | ||
| − | * '''Modification:''' | + | * '''Modification:''' |
* '''Cofactor(s):''' | * '''Cofactor(s):''' | ||
| Line 72: | Line 71: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' | + | * '''Interactions:''' [[DnaA]]-[[YabA]]-[[DnaN]] [http://www.ncbi.nlm.nih.gov/sites/entrez/12060778 PubMed], [[DnaA]]-[[DnaD]] [http://www.ncbi.nlm.nih.gov/sites/entrez/11222620 PubMed] |
| − | |||
| − | |||
| − | * '''Localization:''' cytoplasm [http://www.ncbi.nlm.nih.gov/sites/entrez/ | + | * '''Localization:''' throughout the cytoplasm [http://www.ncbi.nlm.nih.gov/sites/entrez/10844689 PubMed] |
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' |
| − | |||
| − | |||
| − | * '''Swiss prot entry:''' | + | * '''Swiss prot entry:''' |
| − | * '''KEGG entry:''' | + | * '''KEGG entry:''' |
| − | * '''E.C. number:''' | + | * '''E.C. number:''' |
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' ''[[dnaA]]-[[dnaN]]'' [http://www.ncbi.nlm.nih.gov/sites/entrez/2987848 PubMed] |
| − | |||
| − | |||
| − | + | * '''Sigma factor:''' [[SigA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/2987848 PubMed] | |
| − | * ''' | + | * '''Regulation:''' negatively controlled by [[DnaA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/2168872 PubMed] and [[Spo0A]] [http://www.ncbi.nlm.nih.gov/sites/entrez/14651647 PubMed] |
| − | * ''' | + | * '''Regulatory mechanism:''' |
| − | + | * '''Additional information:''' | |
| − | |||
| − | |||
| − | |||
| − | * '''Additional information:''' | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' | + | * '''Mutant:''' |
| − | * '''Expression vector:''' | + | * '''Expression vector:''' |
| − | + | ||
| − | * '''lacZ fusion:''' | + | * '''lacZ fusion:''' |
* '''GFP fusion:''' | * '''GFP fusion:''' | ||
| − | * '''two-hybrid system:''' | + | * '''two-hybrid system:''' |
| − | * '''Antibody:''' | + | * '''Antibody:''' |
=Labs working on this gene/protein= | =Labs working on this gene/protein= | ||
| − | + | [[Philippe Noirot]], Jouy-en-Josas, France [http://locus.jouy.inra.fr/cms/index.php?id=18 homepage] | |
| − | [[ | ||
| − | |||
| − | |||
| − | [http:// | ||
=Your additional remarks= | =Your additional remarks= | ||
| Line 137: | Line 120: | ||
=References= | =References= | ||
| − | # | + | # Imai et al. (2000) Subcellular localization of Dna-initiation proteins of ''Bacillus subtilis'': evidence that chromosome replication begins at either edge of the nucleoids. ''Mol. Microbiol.'' '''36:''' 1037-1048. [http://www.ncbi.nlm.nih.gov/sites/entrez/10844689 PubMed] |
| − | # | + | # Ishigo-Oka et al. (2001) DnaD protein of ''Bacillus subtilis'' interacts with DnaA, the initiator protein of replication. ''J. Bacteriol.'' '''183:''' 2148-2150. [http://www.ncbi.nlm.nih.gov/sites/entrez/11222620 PubMed] |
| − | + | # Noirot-Gros et al. (2002) An expanded view of bacterial DNA replication. ''Proc. Natl. Acad. Sci. USA'' '''99:''' 8342-8347. [http://www.ncbi.nlm.nih.gov/sites/entrez/12060778 PubMed] | |
| − | # | + | # Noirot-Gros et al. (2002) Functional dissection of YabA, a negative regulator of DNA replication initiation in Bacillus Title ''Proc. Natl. Acad. Sci. USA'' '''103:''' 2368-2373. [http://www.ncbi.nlm.nih.gov/sites/entrez/16461910 PubMed] |
| − | + | # Ogasawara et al. (1985) Structure and function of the region of the replication origin of the ''Bacillus subtilis'' chromosome. IV. Transcription of the oriC region and expression of DNA gyrase genes and other open reading frames.''Nucl. Acids Res.'' '''11:''' 2267-2279. [http://www.ncbi.nlm.nih.gov/sites/entrez/2987848 PubMed] | |
| − | # | + | # Fukuoka et al. (1990) Purification and characterization of an initiation protein for chromosomal replication, DnaA, in ''Bacillus subtilis''. ''J. Biochem.'' '''107:''' 732-739. [http://www.ncbi.nlm.nih.gov/sites/entrez/2168872 PubMed] |
| − | + | # Molle et al. (2003) The Spo0A regulon of ''Bacillus subtilis''.''Mol. Microbiol.'' '''50:''' 1683-1701. [http://www.ncbi.nlm.nih.gov/sites/entrez/14651647 PubMed] | |
| − | # | + | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] |
| − | # | ||
| − | |||
| − | # | ||
| − | |||
| − | # | ||
Revision as of 13:16, 11 February 2009
- Description: replication initiation protein
| Gene name | dnaA |
| Synonyms | dnaH, dnaJ, dnaK |
| Essential | yes |
| Product | |
| Function | initiation of chromosome replication |
| MW, pI | 50 kDa, 6.035 |
| Gene length, protein length | 1338 bp, 446 aa |
| Immediate neighbours | |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 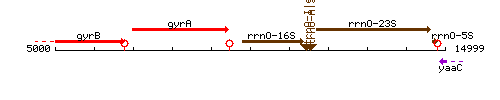
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
essential
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: throughout the cytoplasm PubMed
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry:
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Philippe Noirot, Jouy-en-Josas, France homepage
Your additional remarks
References
- Imai et al. (2000) Subcellular localization of Dna-initiation proteins of Bacillus subtilis: evidence that chromosome replication begins at either edge of the nucleoids. Mol. Microbiol. 36: 1037-1048. PubMed
- Ishigo-Oka et al. (2001) DnaD protein of Bacillus subtilis interacts with DnaA, the initiator protein of replication. J. Bacteriol. 183: 2148-2150. PubMed
- Noirot-Gros et al. (2002) An expanded view of bacterial DNA replication. Proc. Natl. Acad. Sci. USA 99: 8342-8347. PubMed
- Noirot-Gros et al. (2002) Functional dissection of YabA, a negative regulator of DNA replication initiation in Bacillus Title Proc. Natl. Acad. Sci. USA 103: 2368-2373. PubMed
- Ogasawara et al. (1985) Structure and function of the region of the replication origin of the Bacillus subtilis chromosome. IV. Transcription of the oriC region and expression of DNA gyrase genes and other open reading frames.Nucl. Acids Res. 11: 2267-2279. PubMed
- Fukuoka et al. (1990) Purification and characterization of an initiation protein for chromosomal replication, DnaA, in Bacillus subtilis. J. Biochem. 107: 732-739. PubMed
- Molle et al. (2003) The Spo0A regulon of Bacillus subtilis.Mol. Microbiol. 50: 1683-1701. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed