RocG
- Description: trigger enzyme: catabolic glutamate dehydrogenase induced by arginine, ornithine or proline, subject to carbon catabolite repression
| Gene name | rocG |
| Synonyms | |
| Essential | no |
| Product | trigger enzyme: glutamate dehydrogenase (major) |
| Function | arginine utilization, controls the activity of GltC |
| Metabolic function and regulation of this protein in SubtiPathways: Ammonium/ glutamate | |
| MW, pI | 46.2 kDa, 6.28 |
| Gene length, protein length | 1272 bp, 424 amino acids |
| Immediate neighbours | rocA, yweA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 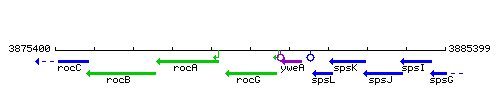
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU37790
Phenotypes of a mutant
Poor growth on complex media such as SP (sporulation medium). No growth in minimal media with arginine as the only carbon source. Rapid accumulation of suppressor mutants (gudB1)
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: L-glutamate + H2O + NAD+ = 2-oxoglutarate + NH3 + NADH (according to Swiss-Prot) L-glutamate + H(2)O + NAD(+) = 2-oxoglutarate + NH(3) + NADH, controls the activity of the GltC transcription activator PubMed
- Protein family: Glu/Leu/Phe/Val dehydrogenases family (according to Swiss-Prot) Glu/Leu/Phe/Val dehydrogenases family
- Paralogous protein(s): GudB
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions: RocG-GltC, this interaction prevents transcription activation of the gltA-gltB operon by GltC PubMed
- Localization:
Database entries
- Structure: 3K92 (super-repressor mutant that is capable of constitutive inactivation of GltC, E93K mutation) PubMed
- UniProt: P39633
- KEGG entry: [3]
- E.C. number: 1.4.1.2
Additional information
Expression and regulation
- Operon: rocG PubMed
- Regulation:
- Regulatory mechanism:
- Additional information:
Activation by RocR requires binding of RocR to a downstream element PubMed
Biological materials
- Mutant: GP747 (spc), GP726 (aphA3), GP810 (tet) available in Stülke lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Stülke lab
- Antibody: available in Stülke lab
Labs working on this gene/protein
Linc Sonenshein, Tufts University, Boston, MA, USA Homepage
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
Enzymatic activity of RocG
Function in the control of GltC activity
Expression of rocG
Structural analysis of glutamate dehydrogenase
Katrin Gunka, Joseph A Newman, Fabian M Commichau, Christina Herzberg, Cecilia Rodrigues, Lorraine Hewitt, Richard J Lewis, Jörg Stülke
Functional dissection of a trigger enzyme: mutations of the bacillus subtilis glutamate dehydrogenase RocG that affect differentially its catalytic activity and regulatory properties.
J Mol Biol: 2010, 400(4);815-27
[PubMed:20630473]
[WorldCat.org]
[DOI]
(I p)
T J Stillman, P J Baker, K L Britton, D W Rice
Conformational flexibility in glutamate dehydrogenase. Role of water in substrate recognition and catalysis.
J Mol Biol: 1993, 234(4);1131-9
[PubMed:8263917]
[WorldCat.org]
[DOI]
(P p)