Difference between revisions of "RibA"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Jstuelk) |
|||
| Line 100: | Line 100: | ||
* '''Operon:''' ''[[ypuE]]-[[ribD]]-[[ribE]]-[[ribA]]-[[ribH]]-[[ribT]]'' [http://www.ncbi.nlm.nih.gov/sites/entrez/8159171 PubMed] | * '''Operon:''' ''[[ypuE]]-[[ribD]]-[[ribE]]-[[ribA]]-[[ribH]]-[[ribT]]'' [http://www.ncbi.nlm.nih.gov/sites/entrez/8159171 PubMed] | ||
| − | * '''[ | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=ribAB_2428389_2429585_-1 ribA] {{PubMed|22383849}} |
| + | |||
| + | * '''Sigma factor:''' [[SigA]] [http://www.ncbi.nlm.nih.gov/sites/entrez/8159171 PubMed] | ||
* '''Regulation:''' expressed in the absence of FMN ([[FMN-box]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/15808508 PubMed] | * '''Regulation:''' expressed in the absence of FMN ([[FMN-box]]) [http://www.ncbi.nlm.nih.gov/sites/entrez/15808508 PubMed] | ||
Revision as of 09:20, 16 April 2012
- Description: GTP cyclohydrolase II/ 3,4-dihydroxy-2-butanone 4-phosphate synthase
| Gene name | ribA |
| Synonyms | |
| Essential | no |
| Product | GTP cyclohydrolase II/ 3,4-dihydroxy-2-butanone 4-phosphate synthase |
| Function | riboflavin biosynthesis |
| Metabolic function and regulation of this protein in SubtiPathways: Riboflavin / FAD | |
| MW, pI | 43 kDa, 5.582 |
| Gene length, protein length | 1194 bp, 398 aa |
| Immediate neighbours | ribH, ribE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 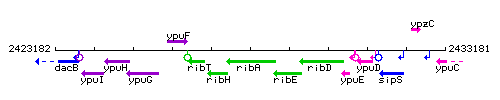
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU23260
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: D-ribulose 5-phosphate = formate + L-3,4-dihydroxybutan-2-one 4-phosphate (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- UniProt: P17620
- KEGG entry: [3]
Additional information
Expression and regulation
- Regulatory mechanism: FMN-box: riboswitch, mediates termination/ antitermination control of the operon, in the absence of FMN: antitermination, in the presence of FMN: termination PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Martin Lehmann, Simone Degen, Hans-Peter Hohmann, Markus Wyss, Adelbert Bacher, Nicholas Schramek
Biosynthesis of riboflavin. Screening for an improved GTP cyclohydrolase II mutant.
FEBS J: 2009, 276(15);4119-29
[PubMed:19583770]
[WorldCat.org]
[DOI]
(I p)
Jingshan Ren, Masayo Kotaka, Michael Lockyer, Heather K Lamb, Alastair R Hawkins, David K Stammers
GTP cyclohydrolase II structure and mechanism.
J Biol Chem: 2005, 280(44);36912-9
[PubMed:16115872]
[WorldCat.org]
[DOI]
(P p)
J Kenneth Wickiser, Wade C Winkler, Ronald R Breaker, Donald M Crothers
The speed of RNA transcription and metabolite binding kinetics operate an FMN riboswitch.
Mol Cell: 2005, 18(1);49-60
[PubMed:15808508]
[WorldCat.org]
[DOI]
(P p)
Wade C Winkler, Smadar Cohen-Chalamish, Ronald R Breaker
An mRNA structure that controls gene expression by binding FMN.
Proc Natl Acad Sci U S A: 2002, 99(25);15908-13
[PubMed:12456892]
[WorldCat.org]
[DOI]
(P p)
V N Mironov, A S Kraev, M L Chikindas, B K Chernov, A I Stepanov, K G Skryabin
Functional organization of the riboflavin biosynthesis operon from Bacillus subtilis SHgw.
Mol Gen Genet: 1994, 242(2);201-8
[PubMed:8159171]
[WorldCat.org]
[DOI]
(P p)
V Azevedo, A Sorokin, S D Ehrlich, P Serror
The transcriptional organization of the Bacillus subtilis 168 chromosome region between the spoVAF and serA genetic loci.
Mol Microbiol: 1993, 10(2);397-405
[PubMed:7934830]
[WorldCat.org]
[DOI]
(P p)
A Sorokin, E Zumstein, V Azevedo, S D Ehrlich, P Serror
The organization of the Bacillus subtilis 168 chromosome region between the spoVA and serA genetic loci, based on sequence data.
Mol Microbiol: 1993, 10(2);385-95
[PubMed:7934829]
[WorldCat.org]
[DOI]
(P p)