Difference between revisions of "RhiN"
| Line 34: | Line 34: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | |||
| − | |||
| − | |||
| − | |||
<br/><br/> | <br/><br/> | ||
| Line 61: | Line 57: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 79: | Line 72: | ||
* '''Kinetic information:''' | * '''Kinetic information:''' | ||
| − | * '''Domains:''' | + | * '''[[Domains]]:''' |
* '''Modification:''' | * '''Modification:''' | ||
| − | * ''' | + | * '''[[Cofactors]]:''' |
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| Line 110: | Line 103: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=yesR_764781_765815_1 yesR] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=yesR_764781_765815_1 yesR] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' | + | * '''[[Sigma factor]]:''' |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 139: | Line 132: | ||
=References= | =References= | ||
| − | <pubmed>,16781735,17449691 </pubmed> | + | <pubmed>,16781735,17449691 24391637 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 13:16, 7 January 2014
- Description: galacturonyl hydrolase, catalyses intracellular degradation of disaccharides generated by YesX
| Gene name | yesR |
| Synonyms | |
| Essential | no |
| Product | galacturonyl hydrolase, catalyses intracellular degradation of disaccharides generated by YesX |
| Function | intracellular degradation of disaccharides generated by YesX |
| Gene expression levels in SubtiExpress: yesR | |
| MW, pI | 38 kDa, 4.757 |
| Gene length, protein length | 1032 bp, 344 aa |
| Immediate neighbours | yesQ, yesS |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 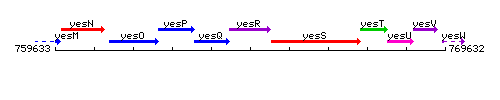
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed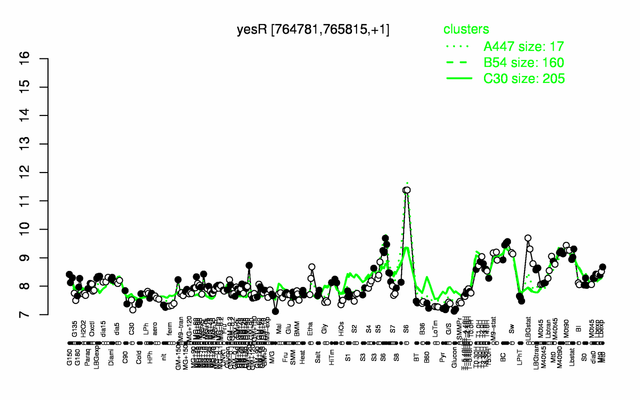
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU07000
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: glycosyl hydrolase 105 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
- Localization:
- cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: O31521
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Regulation:
- induced by pectin PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Irina A Rodionova, Xiaoqing Li, Vera Thiel, Sergey Stolyar, Krista Stanton, James K Fredrickson, Donald A Bryant, Andrei L Osterman, Aaron A Best, Dmitry A Rodionov
Comparative genomics and functional analysis of rhamnose catabolic pathways and regulons in bacteria.
Front Microbiol: 2013, 4;407
[PubMed:24391637]
[WorldCat.org]
[DOI]
(P e)
Akihito Ochiai, Takafumi Itoh, Akiko Kawamata, Wataru Hashimoto, Kousaku Murata
Plant cell wall degradation by saprophytic Bacillus subtilis strains: gene clusters responsible for rhamnogalacturonan depolymerization.
Appl Environ Microbiol: 2007, 73(12);3803-13
[PubMed:17449691]
[WorldCat.org]
[DOI]
(P p)
Takafumi Itoh, Akihito Ochiai, Bunzo Mikami, Wataru Hashimoto, Kousaku Murata
A novel glycoside hydrolase family 105: the structure of family 105 unsaturated rhamnogalacturonyl hydrolase complexed with a disaccharide in comparison with family 88 enzyme complexed with the disaccharide.
J Mol Biol: 2006, 360(3);573-85
[PubMed:16781735]
[WorldCat.org]
[DOI]
(P p)