Difference between revisions of "RadC"
| Line 88: | Line 88: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' | + | * '''Operon:''' ''radC'' (according to [http://dbtbs.hgc.jp/COG/prom/radC.html DBTBS]) |
| − | * '''[[Sigma factor]]:''' [[SigM]] | + | * '''[[Sigma factor]]:''' [[SigM]] {{PubMed|12775685}} |
* '''Regulation:''' | * '''Regulation:''' | ||
| + | ** expressed in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress ([[SigM]]) {{PubMed|12775685}} | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| Line 118: | Line 119: | ||
=References= | =References= | ||
| − | <pubmed>17434969,,16479537 </pubmed> | + | <pubmed>17434969,,16479537 12775685 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 20:32, 13 December 2009
- Description: probable DNA repair protein
| Gene name | radC |
| Synonyms | ysxA |
| Essential | no |
| Product | probable DNA repair protein |
| Function | DNA repair |
| MW, pI | 25 kDa, 7.829 |
| Gene length, protein length | 693 bp, 231 aa |
| Immediate neighbours | mreB, maf |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 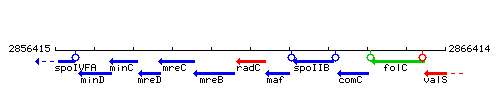
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU28040
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: radC family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: Cytoplasm (Homogeneous) PubMed
Database entries
- Structure:
- UniProt: Q02170
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: radC (according to DBTBS)
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Adrian J Jervis, Penny D Thackray, Chris W Houston, Malcolm J Horsburgh, Anne Moir
SigM-responsive genes of Bacillus subtilis and their promoters.
J Bacteriol: 2007, 189(12);4534-8
[PubMed:17434969]
[WorldCat.org]
[DOI]
(P p)
Jean-Christophe Meile, Ling Juan Wu, S Dusko Ehrlich, Jeff Errington, Philippe Noirot
Systematic localisation of proteins fused to the green fluorescent protein in Bacillus subtilis: identification of new proteins at the DNA replication factory.
Proteomics: 2006, 6(7);2135-46
[PubMed:16479537]
[WorldCat.org]
[DOI]
(P p)
Penny D Thackray, Anne Moir
SigM, an extracytoplasmic function sigma factor of Bacillus subtilis, is activated in response to cell wall antibiotics, ethanol, heat, acid, and superoxide stress.
J Bacteriol: 2003, 185(12);3491-8
[PubMed:12775685]
[WorldCat.org]
[DOI]
(P p)