PyrB
- Description: aspartate carbamoyltransferase
| Gene name | pyrB |
| Synonyms | |
| Essential | no |
| Product | aspartate carbamoyltransferase |
| Function | pyrimidine biosynthesis |
| Metabolic function and regulation of this protein in SubtiPathways: Pyrimidines, Nucleotides (regulation) | |
| MW, pI | 34 kDa, 5.341 |
| Gene length, protein length | 912 bp, 304 aa |
| Immediate neighbours | pyrP, pyrC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 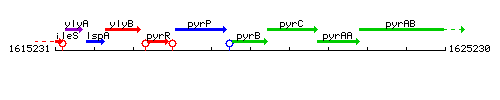
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU15490
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Carbamoyl phosphate + L-aspartate = phosphate + N-carbamoyl-L-aspartate (according to Swiss-Prot)
- Protein family: ATCase/OTCase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation on Ser-303 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure: 2AT2
- Swiss prot entry: P05654
- KEGG entry: [3]
- E.C. number: 2.1.3.2
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulatory mechanism:
- PyrR: RNA switch, transcription termination/ antitermination (in the presence of uridine nucleotides: termination, in their absence: antitermination) PubMed
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Boris Macek, Ivan Mijakovic, Jesper V Olsen, Florian Gnad, Chanchal Kumar, Peter R Jensen, Matthias Mann
The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis.
Mol Cell Proteomics: 2007, 6(4);697-707
[PubMed:17218307]
[WorldCat.org]
[DOI]
(P p)
Hesheng Zhang, Robert L Switzer
Transcriptional pausing in the Bacillus subtilis pyr operon in vitro: a role in transcriptional attenuation?
J Bacteriol: 2003, 185(16);4764-71
[PubMed:12896995]
[WorldCat.org]
[DOI]
(P p)
Y Lu, R J Turner, R L Switzer
Roles of the three transcriptional attenuators of the Bacillus subtilis pyrimidine biosynthetic operon in the regulation of its expression.
J Bacteriol: 1995, 177(5);1315-25
[PubMed:7868607]
[WorldCat.org]
[DOI]
(P p)
P Hu, R L Switzer
Evidence for substrate stabilization in regulation of the degradation of Bacillus subtilis aspartate transcarbamylase in vivo.
Arch Biochem Biophys: 1995, 316(1);260-6
[PubMed:7840626]
[WorldCat.org]
[DOI]
(P p)
R J Turner, Y Lu, R L Switzer
Regulation of the Bacillus subtilis pyrimidine biosynthetic (pyr) gene cluster by an autogenous transcriptional attenuation mechanism.
J Bacteriol: 1994, 176(12);3708-22
[PubMed:8206849]
[WorldCat.org]
[DOI]
(P p)
C L Quinn, B T Stephenson, R L Switzer
Functional organization and nucleotide sequence of the Bacillus subtilis pyrimidine biosynthetic operon.
J Biol Chem: 1991, 266(14);9113-27
[PubMed:1709162]
[WorldCat.org]
(P p)
C G Lerner, R L Switzer
Cloning and structure of the Bacillus subtilis aspartate transcarbamylase gene (pyrB).
J Biol Chem: 1986, 261(24);11156-65
[PubMed:3015959]
[WorldCat.org]
(P p)
J S Brabson, M R Maurizi, R L Switzer
Aspartate transcarbamylase from Bacillus subtilis.
Methods Enzymol: 1985, 113;627-35
[PubMed:3937019]
[WorldCat.org]
[DOI]
(P p)
R W Bond, R L Switzer
Degradation of aspartate transcarbamylase in Bacillus subtilis is deficient in rel mutants but is not mediated by guanosine polyphosphates.
J Bacteriol: 1984, 158(2);746-8
[PubMed:6427186]
[WorldCat.org]
[DOI]
(P p)