Difference between revisions of "PurA"
| Line 14: | Line 14: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || purine biosynthesis | |style="background:#ABCDEF;" align="center"|'''Function''' || purine biosynthesis | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/gene_regulation_nucleotides.html Nucleotides (regulation)]''' | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/purine_biosynthesis.html Purine synthesis], [http://subtiwiki.uni-goettingen.de/pathways/gene_regulation_nucleotides.html Nucleotides (regulation)]''' |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 47 kDa, 5.459 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 47 kDa, 5.459 | ||
Revision as of 12:56, 18 June 2009
- Description: adenylosuccinate synthetase
| Gene name | purA |
| Synonyms | |
| Essential | no |
| Product | adenylosuccinate synthetase |
| Function | purine biosynthesis |
| Metabolic function and regulation of this protein in SubtiPathways: Purine synthesis, Nucleotides (regulation) | |
| MW, pI | 47 kDa, 5.459 |
| Gene length, protein length | 1290 bp, 430 aa |
| Immediate neighbours | trnY-Lys, yycE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 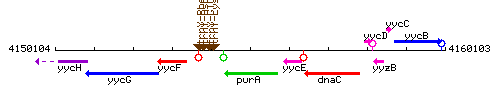
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU40420
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: GTP + IMP + L-aspartate = GDP + phosphate + N(6)-(1,2-dicarboxyethyl)-AMP (according to Swiss-Prot)
- Protein family: adenylosuccinate synthetase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: P29726
- KEGG entry: [3]
- E.C. number: 6.3.4.4
Additional information
Expression and regulation
- Operon:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
M Weng, P L Nagy, H Zalkin
Identification of the Bacillus subtilis pur operon repressor.
Proc Natl Acad Sci U S A: 1995, 92(16);7455-9
[PubMed:7638212]
[WorldCat.org]
[DOI]
(P p)
H H Saxild, P Nygaard
Regulation of levels of purine biosynthetic enzymes in Bacillus subtilis: effects of changing purine nucleotide pools.
J Gen Microbiol: 1991, 137(10);2387-94
[PubMed:1722815]
[WorldCat.org]
[DOI]
(P p)