PtsG
- Description: trigger enzyme: major glucose permease of the PTS, EIICBA(Glc)
| Gene name | ptsG |
| Synonyms | ptsX, crr |
| Essential | no |
| Product | glucose-specific enzyme IICBA component |
| Function | glucose transport and phosphorylation, control of GlcT activity |
| MW, pI | 75,3 kDa, 5.40 |
| Gene length, protein length | 2097 bp, 699 amino acids |
| Immediate neighbours | glcT, ptsH |
| Gene sequence (+200bp) | Protein sequence |
| Caution: The sequence for this gene in SubtiList contains errors | |
Genetic context 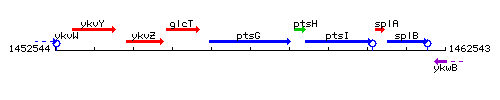
| |
Contents
The gene
Basic information
- Coordinates: 1456496 - 1458592
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transport and phosphorylation of glucose, receives a phosphate from HPr at the IIA domain (His-620), the phosphate group is then transferred to the IIB domain (Cys-461) an finally to the incoming glucose. In the absence of glucose, PtsG phosphorylates and thereby inactivates the transcriptional antiterminator GlcT.
- Protein family: PTS enzyme II, glucose family
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- 11x transmembrane domain (16–36, 89–109, 139–159, 180–200, 233–253, 283–303, 313–333, 338–358, 365–385, 388–408)
- PTS EIIC domain ( 1-424)
- PTS EIIB domain (439–520)
- PTS EIIA domain (568–672)
- Modification: transient phosphorylation (HPr-dependent) on His-620, then internal phosphotransfer from His-620 to Cys-461
- Cofactor(s):
- Effectors of protein activity:
- Localization: membrane protein NCBI
Database entries
- Swiss prot entry: [3]
- KEGG entry: [4]
- E.C. number: [5]
Additional information
Expression and regulation
- Regulation: induction by glucose
- Regulatory mechanism: transcriptional antitermination via the GlcT-dependent RNA-switch PubMed
- Additional information:
Biological materials
- Mutant: GP474 (cat), QB5436 (spc), QB5445 (erm), available in Stülke lab
- Expression vector: pGP123 (domains BA, in pWH844), pGP123 (domains BA, mut: H620D, in pWH844), pGP428 (EIIB, in pWH844), pGP437(EIIA in pWH844, with thrombin cleavage site), available in Stülke lab
- lacZ fusion: pGP34 (pAC5), pGP66 (pAC7), pGP606 (mutant terminator, pAC6), pGP532 (pAC7), series of promoter deletions are available in pAC5 and pAC6, series of RAT mutants are available in pAC6, available in Stülke lab
- GFP fusion:
- Antibody:
Labs working on this gene/protein
Jörg Stülke, University of Göttingen, Germany Homepage
Your additional remarks
References
- Stülke J, Martin-Verstraete I, Zagorec M (1997) Induction of the Bacillus subtilis ptsGHI operon by glucose is controlled by a novel antiterminator, GlcT Mol Microbiol. 25: 65-78. PubMed
- Bachem S, Stülke J. (1998) Regulation of the Bacillus subtilis GlcT antiterminator protein by components of the phosphotransferase system. J Bacteriol. 180: 5319-26 PubMed
- Bachem, S., Faires, N., & Stülke, J. (1997) Characterization of the presumptive phosphorylation sites of the Bacillus subtilis glucose permease by site-directed mutagenesis: Implication in glucose transport and catabolite repression. FEMS Microbiol. L. 156: 233-238. PubMed
- Gonzy-Tréboul, G., de Waard, J. H., Zagorec, M., and Postma, P.W. (1991). The glucose permease of the phosphotransferase system of Bacillus subtilis: Evidence for IIGlc and IIIGlc domains. Mol. Microbiol. 5, 1241-1249. PubMed
- Langbein, I., Bachem, S. & Stülke, J. (1999) Specific interaction of the RNA binding domain of the Bacillus subtilis transcriptional antiterminator GlcT with its RNA target, RAT. J. Mol. Biol. 293: 795-805. PubMed
- Schilling, O., Herzberg, C., Hertrich, T., Vörsmann, H., Jessen, D., Hübner, S., Titgemeyer, F. & Stülke, J. (2006) Keeping signals straight in transcription regulation: specificity determinants for the interaction of a family of conserved bacterial RNA-protein couples. Nucl. Acids Res. 34: 6102-6115. PubMed
- Schilling, O., Langbein, I., Müller, M., Schmalisch, M. & Stülke, J. (2004) A protein-dependent riboswitch controlling ptsGHI operon expression in Bacillus subtilis: RNA structure rather than sequence provides interaction specificity. Nucl. Acids Res. 32: 2853-2864. PubMed
- Schmalisch, M., Bachem, S. & Stülke, J. (2003) Control of the Bacillus subtilis antiterminator protein GlcT by phosphorylation: Elucidation of the phosphorylation chain leading to inactivation of GlcT. J. Biol. Chem. 278: 51108-51115. PubMed
- Zagorec, M. & Postma, P. (1992). Cloning and nucleotide sequence of the ptsG gene of Bacillus subtilis. Mol Gen Genet 234, 325-328. PubMed
- Sutrina, S. L., Reddy, P., Saier, M. H., Jr & Reizer, J. (1990). The glucose permease of Bacillus subtilis is a single polypeptide chain that functions to energize the sucrose permease. J Biol Chem 265, 18581-18589. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed