Difference between revisions of "PsdS"
| Line 30: | Line 30: | ||
<br/><br/> | <br/><br/> | ||
| + | |||
| + | = [[Categories]] containing this gene/protein = | ||
| + | {{SubtiWiki category|[[protein modification]]}}, | ||
| + | {{SubtiWiki category|[[transcription factors and their control]]}}, | ||
| + | {{SubtiWiki category|[[resistance against toxins/ antibiotics]]}}, | ||
| + | {{SubtiWiki category|[[membrane proteins]]}}, | ||
| + | {{SubtiWiki category|[[phosphoproteins]]}} | ||
| + | |||
| + | = This gene is a member of the following [[regulons]] = | ||
| + | |||
=The gene= | =The gene= | ||
| Line 49: | Line 59: | ||
| − | + | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
Revision as of 13:24, 9 December 2010
- Description: two-component sensor kinase, control of psdA-psdB in response to lipid II-binding lantibiotics, such as nisin and gallidermin
| Gene name | psdS |
| Synonyms | yvcQ |
| Essential | no |
| Product | two-component sensor kinase |
| Function | resistance against toxic peptides |
| MW, pI | 40 kDa, 7.358 |
| Gene length, protein length | 1068 bp, 356 aa |
| Immediate neighbours | psdA, psdR |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 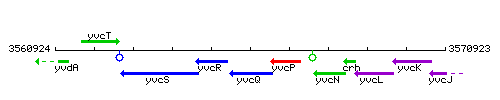
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
protein modification, transcription factors and their control, resistance against toxins/ antibiotics, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU34710
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: autophosphorylation, phosphorylation of PsdR
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains: two transmembrane segments, C-terminal histidine phosphotransferase domain
- Modification: autophosphorylation on a His residue
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: O06979
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Anna Staroń, Dora Elisabeth Finkeisen, Thorsten Mascher
Peptide antibiotic sensing and detoxification modules of Bacillus subtilis.
Antimicrob Agents Chemother: 2011, 55(2);515-25
[PubMed:21078927]
[WorldCat.org]
[DOI]
(I p)
Eva Rietkötter, Diana Hoyer, Thorsten Mascher
Bacitracin sensing in Bacillus subtilis.
Mol Microbiol: 2008, 68(3);768-85
[PubMed:18394148]
[WorldCat.org]
[DOI]
(I p)
Thorsten Mascher, Neil G Margulis, Tao Wang, Rick W Ye, John D Helmann
Cell wall stress responses in Bacillus subtilis: the regulatory network of the bacitracin stimulon.
Mol Microbiol: 2003, 50(5);1591-604
[PubMed:14651641]
[WorldCat.org]
[DOI]
(P p)
C Fabret, V A Feher, J A Hoch
Two-component signal transduction in Bacillus subtilis: how one organism sees its world.
J Bacteriol: 1999, 181(7);1975-83
[PubMed:10094672]
[WorldCat.org]
[DOI]
(P p)