Difference between revisions of "PonA"
(→Categories containing this gene/protein) |
(→Extended information on the protein) |
||
| Line 82: | Line 82: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' | + | * '''[[SubtInteract|Interactions]]:''' |
| − | * '''Localization:''' | + | * '''[[Localization]]:''' |
** membrane associated [http://www.ncbi.nlm.nih.gov/pubmed/18763711 PubMed] | ** membrane associated [http://www.ncbi.nlm.nih.gov/pubmed/18763711 PubMed] | ||
** localizes to the spore septum during sporulation {{PubMed|15758244,15262952}} | ** localizes to the spore septum during sporulation {{PubMed|15758244,15262952}} | ||
Revision as of 10:16, 2 November 2011
- Description: penicillin-binding protein 1A/1B
| Gene name | ponA |
| Synonyms | |
| Essential | no |
| Product | penicillin-binding protein 1A/1B |
| Function | bifunctional glucosyltransferase/ transpeptidase |
| MW, pI | 99 kDa, 4.752 |
| Gene length, protein length | 2742 bp, 914 aa |
| Immediate neighbours | recU, ypoC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 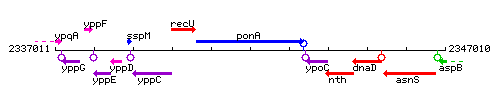
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
cell wall synthesis, cell envelope stress proteins (controlled by SigM, V, W, X, Y), membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU22320
Phenotypes of a mutant
- prevents bulging of the cells when grown at low Mg(2+) concentrations, suppresses the lethal effect of a mreB mutation PubMed
- deletion of ponA restores growth and normal shape of a yvcK mutant on gluconeogenic carbon sources PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P39793
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation: constitutive during vegetative growth PubMed
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jeff Errington lab
- Antibody:
Labs working on this gene/protein
Jeff Errington, Newcastle University, UK homepage
Your additional remarks
References
Reviews
Original publications
Elodie Foulquier, Frédérique Pompeo, Alain Bernadac, Leon Espinosa, Anne Galinier
The YvcK protein is required for morphogenesis via localization of PBP1 under gluconeogenic growth conditions in Bacillus subtilis.
Mol Microbiol: 2011, 80(2);309-18
[PubMed:21320184]
[WorldCat.org]
[DOI]
(I p)
Yoshikazu Kawai, Richard A Daniel, Jeffery Errington
Regulation of cell wall morphogenesis in Bacillus subtilis by recruitment of PBP1 to the MreB helix.
Mol Microbiol: 2009, 71(5);1131-44
[PubMed:19192185]
[WorldCat.org]
[DOI]
(I p)
Hannes Hahne, Susanne Wolff, Michael Hecker, Dörte Becher
From complementarity to comprehensiveness--targeting the membrane proteome of growing Bacillus subtilis by divergent approaches.
Proteomics: 2008, 8(19);4123-36
[PubMed:18763711]
[WorldCat.org]
[DOI]
(I p)
Dennis Claessen, Robyn Emmins, Leendert W Hamoen, Richard A Daniel, Jeff Errington, David H Edwards
Control of the cell elongation-division cycle by shuttling of PBP1 protein in Bacillus subtilis.
Mol Microbiol: 2008, 68(4);1029-46
[PubMed:18363795]
[WorldCat.org]
[DOI]
(I p)
Warawan Eiamphungporn, John D Helmann
The Bacillus subtilis sigma(M) regulon and its contribution to cell envelope stress responses.
Mol Microbiol: 2008, 67(4);830-48
[PubMed:18179421]
[WorldCat.org]
[DOI]
(P p)
Dirk-Jan Scheffers, Jeffery Errington
PBP1 is a component of the Bacillus subtilis cell division machinery.
J Bacteriol: 2004, 186(15);5153-6
[PubMed:15262952]
[WorldCat.org]
[DOI]
(P p)
Dirk-Jan Scheffers, Laura J F Jones, Jeffery Errington
Several distinct localization patterns for penicillin-binding proteins in Bacillus subtilis.
Mol Microbiol: 2004, 51(3);749-64
[PubMed:14731276]
[WorldCat.org]
[DOI]
(P p)
L B Pedersen, E R Angert, P Setlow
Septal localization of penicillin-binding protein 1 in Bacillus subtilis.
J Bacteriol: 1999, 181(10);3201-11
[PubMed:10322023]
[WorldCat.org]
[DOI]
(P p)
T Murray, D L Popham, P Setlow
Bacillus subtilis cells lacking penicillin-binding protein 1 require increased levels of divalent cations for growth.
J Bacteriol: 1998, 180(17);4555-63
[PubMed:9721295]
[WorldCat.org]
[DOI]
(P p)
D L Popham, P Setlow
Phenotypes of Bacillus subtilis mutants lacking multiple class A high-molecular-weight penicillin-binding proteins.
J Bacteriol: 1996, 178(7);2079-85
[PubMed:8606187]
[WorldCat.org]
[DOI]
(P p)
D L Popham, P Setlow
Cloning, nucleotide sequence, and mutagenesis of the Bacillus subtilis ponA operon, which codes for penicillin-binding protein (PBP) 1 and a PBP-related factor.
J Bacteriol: 1995, 177(2);326-35
[PubMed:7814321]
[WorldCat.org]
[DOI]
(P p)