Difference between revisions of "PgcA"
Raphael2215 (talk | contribs) |
|||
| Line 13: | Line 13: | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || interconversion of glucose 6-phosphate and alpha-glucose 1-phosphate | |style="background:#ABCDEF;" align="center"|'''Function''' || interconversion of glucose 6-phosphate and alpha-glucose 1-phosphate | ||
|- | |- | ||
| − | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http:// | + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://subtiwiki.uni-goettingen.de/apps/expression/ ''Subti''Express]''': [http://subtiwiki.uni-goettingen.de/apps/expression/expression.php?search=BSU09310 pgcA] |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/fatty_acid_synthesis.html Lipid synthesis]''' | |colspan="2" style="background:#FAF8CC;" align="center"| '''Metabolic function and regulation of this protein in [[SubtiPathways|''Subti''Pathways]]: <br/>[http://subtiwiki.uni-goettingen.de/pathways/fatty_acid_synthesis.html Lipid synthesis]''' | ||
| Line 23: | Line 23: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[glpD]]'', ''[[yhcY]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[glpD]]'', ''[[yhcY]]'' | ||
|- | |- | ||
| − | | | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU09310 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU09310 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU09310 Advanced_DNA] |
|- | |- | ||
|- | |- | ||
Revision as of 12:25, 13 May 2013
- Description: alpha-phosphoglucomutase, required for UDP-glucose synthesis, inhibits FtsZ ring assembly (indirect effect due to a defect in UDP-glucose synthesis)
| Gene name | pgcA |
| Synonyms | yhxB, gtaC, gtaE |
| Essential | no |
| Product | alpha-phosphoglucomutase |
| Function | interconversion of glucose 6-phosphate and alpha-glucose 1-phosphate |
| Gene expression levels in SubtiExpress: pgcA | |
| Metabolic function and regulation of this protein in SubtiPathways: Lipid synthesis | |
| MW, pI | 62 kDa, 4.913 |
| Gene length, protein length | 1695 bp, 565 aa |
| Immediate neighbours | glpD, yhcY |
| Sequences | Protein DNA Advanced_DNA |
Genetic context 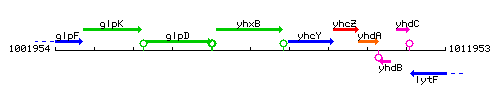
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed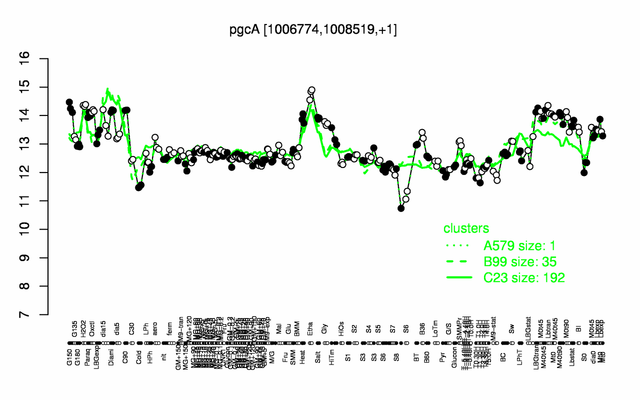
| |
Contents
Categories containing this gene/protein
cell wall synthesis, lipid metabolism/ other, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU09310
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Alpha-D-glucose 1-phosphate = alpha-D-glucose 6-phosphate (according to Swiss-Prot)
- Protein family: phosphohexose mutase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P18159
- KEGG entry: [2]
- E.C. number: 5.4.2.2
Additional information
PgcA inhibits FtsZ ring assembly (indirect effect due to a defect in UDP-glucose synthesis)PubMed
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Massimiliano Marvasi, Pieter T Visscher, Lilliam Casillas Martinez
Exopolymeric substances (EPS) from Bacillus subtilis: polymers and genes encoding their synthesis.
FEMS Microbiol Lett: 2010, 313(1);1-9
[PubMed:20735481]
[WorldCat.org]
[DOI]
(I p)
Original publications
Additional publications: PubMed
Christine Eymann, Dörte Becher, Jörg Bernhardt, Katrin Gronau, Anja Klutzny, Michael Hecker
Dynamics of protein phosphorylation on Ser/Thr/Tyr in Bacillus subtilis.
Proteomics: 2007, 7(19);3509-26
[PubMed:17726680]
[WorldCat.org]
[DOI]
(P p)
Richard B Weart, Amy H Lee, An-Chun Chien, Daniel P Haeusser, Norbert S Hill, Petra Anne Levin
A metabolic sensor governing cell size in bacteria.
Cell: 2007, 130(2);335-47
[PubMed:17662947]
[WorldCat.org]
[DOI]
(P p)
Alain Lévine, Françoise Vannier, Cédric Absalon, Lauriane Kuhn, Peter Jackson, Elaine Scrivener, Valérie Labas, Joëlle Vinh, Patrick Courtney, Jérôme Garin, Simone J Séror
Analysis of the dynamic Bacillus subtilis Ser/Thr/Tyr phosphoproteome implicated in a wide variety of cellular processes.
Proteomics: 2006, 6(7);2157-73
[PubMed:16493705]
[WorldCat.org]
[DOI]
(P p)
Vladimir Lazarevic, Blazenka Soldo, Noël Médico, Harold Pooley, Sierd Bron, Dimitri Karamata
Bacillus subtilis alpha-phosphoglucomutase is required for normal cell morphology and biofilm formation.
Appl Environ Microbiol: 2005, 71(1);39-45
[PubMed:15640167]
[WorldCat.org]
[DOI]
(P p)
Steven S Branda, José Eduardo González-Pastor, Etienne Dervyn, S Dusko Ehrlich, Richard Losick, Roberto Kolter
Genes involved in formation of structured multicellular communities by Bacillus subtilis.
J Bacteriol: 2004, 186(12);3970-9
[PubMed:15175311]
[WorldCat.org]
[DOI]
(P p)