Difference between revisions of "PcrA"
| Line 133: | Line 133: | ||
* '''Additional information:''' | * '''Additional information:''' | ||
| + | ** number of protein molecules per cell (minimal medium with glucose and ammonium): 114 {{PubMed|24696501}} | ||
| + | ** number of protein molecules per cell (complex medium with amino acids, without glucose): 250 {{PubMed|24696501}} | ||
=Biological materials = | =Biological materials = | ||
Revision as of 09:36, 17 April 2014
- Description: ATP-dependent DNA helicase, facilitates unwinding of ICEBs1 DNA for horizontal transfer
| Gene name | pcrA |
| Synonyms | yerF |
| Essential | yes PubMed |
| Product | ATP-dependent DNA helicase |
| Function | plasmid rolling-circle replication, conjugative transfer of ICEBs1 |
| Gene expression levels in SubtiExpress: pcrA | |
| Interactions involving this protein in SubtInteract: PcrA | |
| MW, pI | 83 kDa, 5.767 |
| Gene length, protein length | 2217 bp, 739 aa |
| Immediate neighbours | pcrB, ligA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 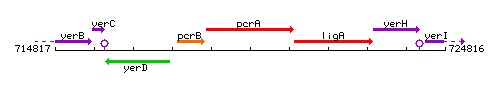
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed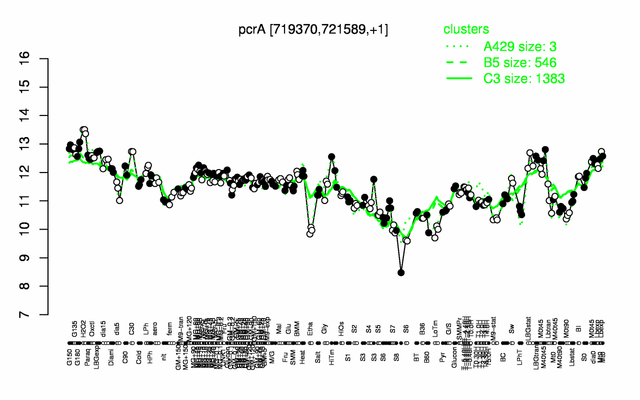
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, mobile genetic elements, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU06610
Phenotypes of a mutant
essential PubMed
Database entries
- BsubCyc: BSU06610
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: uvrD-like helicase C-terminal domain (according to Swiss-Prot)
- Paralogous protein(s): YjcD
Extended information on the protein
- Kinetic information:
- Domains:
- two RecA-like domains PubMed
- intrinxically disordered C-terminal domain (for interaction with RNA polymerase) PubMed
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- unwinding of the double-stranded ICEBs1 DNA is stimulated by HelP PubMed
- helicase activity is stimulated by interaction with RNA polymerase PubMed
Database entries
- BsubCyc: BSU06610
- Structure:
- UniProt: O34580
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Elisabeth Grohmann
Autonomous plasmid-like replication of Bacillus ICEBs1: a general feature of integrative conjugative elements?
Mol Microbiol: 2010, 75(2);261-3
[PubMed:19943906]
[WorldCat.org]
[DOI]
(I p)
Original Publications
Emma J Gwynn, Abigail J Smith, Colin P Guy, Nigel J Savery, Peter McGlynn, Mark S Dillingham
The conserved C-terminus of the PcrA/UvrD helicase interacts directly with RNA polymerase.
PLoS One: 2013, 8(10);e78141
[PubMed:24147116]
[WorldCat.org]
[DOI]
(I e)
Jacob Thomas, Catherine A Lee, Alan D Grossman
A conserved helicase processivity factor is needed for conjugation and replication of an integrative and conjugative element.
PLoS Genet: 2013, 9(1);e1003198
[PubMed:23326247]
[WorldCat.org]
[DOI]
(I p)
Olivier Delumeau, François Lecointe, Jan Muntel, Alain Guillot, Eric Guédon, Véronique Monnet, Michael Hecker, Dörte Becher, Patrice Polard, Philippe Noirot
The dynamic protein partnership of RNA polymerase in Bacillus subtilis.
Proteomics: 2011, 11(15);2992-3001
[PubMed:21710567]
[WorldCat.org]
[DOI]
(I p)
Jeehae Park, Sua Myong, Anita Niedziela-Majka, Kyung Suk Lee, Jin Yu, Timothy M Lohman, Taekjip Ha
PcrA helicase dismantles RecA filaments by reeling in DNA in uniform steps.
Cell: 2010, 142(4);544-55
[PubMed:20723756]
[WorldCat.org]
[DOI]
(I p)
Catherine A Lee, Ana Babic, Alan D Grossman
Autonomous plasmid-like replication of a conjugative transposon.
Mol Microbiol: 2010, 75(2);268-79
[PubMed:19943900]
[WorldCat.org]
[DOI]
(I p)
Ye Yang, Shuo-Xing Dou, Hua Ren, Peng-Ye Wang, Xing-Dong Zhang, Min Qian, Bing-Yi Pan, Xu Guang Xi
Evidence for a functional dimeric form of the PcrA helicase in DNA unwinding.
Nucleic Acids Res: 2008, 36(6);1976-89
[PubMed:18276648]
[WorldCat.org]
[DOI]
(I p)
Syam P Anand, Haocheng Zheng, Piero R Bianco, Sanford H Leuba, Saleem A Khan
DNA helicase activity of PcrA is not required for the displacement of RecA protein from DNA or inhibition of RecA-mediated strand exchange.
J Bacteriol: 2007, 189(12);4502-9
[PubMed:17449621]
[WorldCat.org]
[DOI]
(P p)
Nora Au, Elke Kuester-Schoeck, Veena Mandava, Laura E Bothwell, Susan P Canny, Karen Chachu, Sierra A Colavito, Shakierah N Fuller, Eli S Groban, Laura A Hensley, Theresa C O'Brien, Amish Shah, Jessica T Tierney, Louise L Tomm, Thomas M O'Gara, Alexi I Goranov, Alan D Grossman, Charles M Lovett
Genetic composition of the Bacillus subtilis SOS system.
J Bacteriol: 2005, 187(22);7655-66
[PubMed:16267290]
[WorldCat.org]
[DOI]
(P p)
M-F Noirot-Gros, P Soultanas, D B Wigley, S D Ehrlich, P Noirot, M-A Petit
The beta-propeller protein YxaL increases the processivity of the PcrA helicase.
Mol Genet Genomics: 2002, 267(3);391-400
[PubMed:12073041]
[WorldCat.org]
[DOI]
(P p)
P Soultanas, M S Dillingham, P Wiley, M R Webb, D B Wigley
Uncoupling DNA translocation and helicase activity in PcrA: direct evidence for an active mechanism.
EMBO J: 2000, 19(14);3799-810
[PubMed:10899133]
[WorldCat.org]
[DOI]
(P p)
S S Velankar, P Soultanas, M S Dillingham, H S Subramanya, D B Wigley
Crystal structures of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism.
Cell: 1999, 97(1);75-84
[PubMed:10199404]
[WorldCat.org]
[DOI]
(P p)
M A Petit, E Dervyn, M Rose, K D Entian, S McGovern, S D Ehrlich, C Bruand
PcrA is an essential DNA helicase of Bacillus subtilis fulfilling functions both in repair and rolling-circle replication.
Mol Microbiol: 1998, 29(1);261-73
[PubMed:9701819]
[WorldCat.org]
[DOI]
(P p)