Difference between revisions of "PbpA"
| Line 26: | Line 26: | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:pbpA_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:pbpA_context.gif]] | ||
<div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | <div align="right"> <small>This image was kindly provided by [http://genolist.pasteur.fr/SubtiList/ SubtiList]</small></div> | ||
| + | |- | ||
| + | |colspan="2" |'''[http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=pbpA_2581771_2583921_-1 Expression at a glance]'''   {{PubMed|22383849}}<br/>[[Image:pbpA_expression.png|500px]] | ||
|- | |- | ||
|} | |} | ||
__TOC__ | __TOC__ | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | <br/><br/><br/><br/> | ||
| + | |||
<br/><br/> | <br/><br/> | ||
Revision as of 08:33, 20 April 2012
- Description: penicillin-binding protein PBP 2A
| Gene name | pbpA |
| Synonyms | yqgF |
| Essential | no |
| Product | penicillin-binding protein PBP 2A |
| Function | formation of a rod-shaped peptidoglycan cell wall, spore outgrowth |
| Interactions involving this protein in SubtInteract: PbpA | |
| MW, pI | 79 kDa, 9.571 |
| Gene length, protein length | 2148 bp, 716 aa |
| Immediate neighbours | pstS, yqgE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 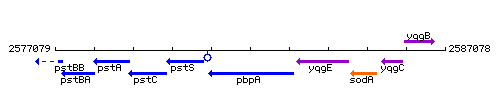
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed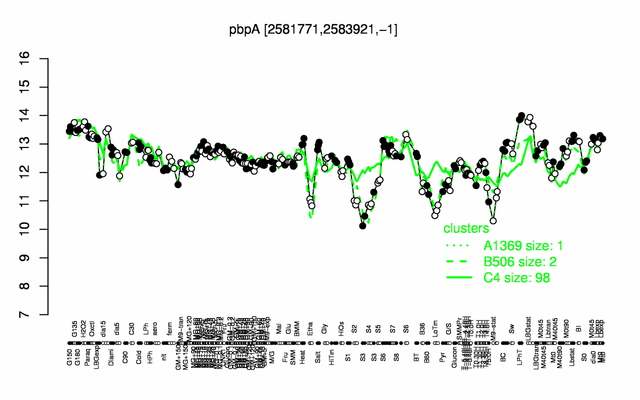
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU25000
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- part of the cell wall biosynthetic complex PubMed
- folding requires PrsA PubMed
- Localization: extracellular (signal peptide) PubMed
Database entries
- Structure:
- UniProt: P54488
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Jeff Errington lab
- Antibody:
Labs working on this gene/protein
Jeff Errington, Newcastle University, UK homepage
Your additional remarks
References
Lehnik-Habrink M, Schaffer M, Mäder U, Diethmaier C, Herzberg C, Stülke J RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y. Mol Microbiol. 2011 81(6): 1459-1473. PubMed:21815947
Ethan C Garner, Remi Bernard, Wenqin Wang, Xiaowei Zhuang, David Z Rudner, Tim Mitchison
Coupled, circumferential motions of the cell wall synthesis machinery and MreB filaments in B. subtilis.
Science: 2011, 333(6039);222-5
[PubMed:21636745]
[WorldCat.org]
[DOI]
(I p)
Julia Domínguez-Escobar, Arnaud Chastanet, Alvaro H Crevenna, Vincent Fromion, Roland Wedlich-Söldner, Rut Carballido-López
Processive movement of MreB-associated cell wall biosynthetic complexes in bacteria.
Science: 2011, 333(6039);225-8
[PubMed:21636744]
[WorldCat.org]
[DOI]
(I p)
Hanne-Leena Hyyryläinen, Bogumila C Marciniak, Kathleen Dahncke, Milla Pietiäinen, Pascal Courtin, Marika Vitikainen, Raili Seppala, Andreas Otto, Dörte Becher, Marie-Pierre Chapot-Chartier, Oscar P Kuipers, Vesa P Kontinen
Penicillin-binding protein folding is dependent on the PrsA peptidyl-prolyl cis-trans isomerase in Bacillus subtilis.
Mol Microbiol: 2010, 77(1);108-27
[PubMed:20487272]
[WorldCat.org]
[DOI]
(I p)
Birgit Voigt, Haike Antelmann, Dirk Albrecht, Armin Ehrenreich, Karl-Heinz Maurer, Stefan Evers, Gerhard Gottschalk, Jan Maarten van Dijl, Thomas Schweder, Michael Hecker
Cell physiology and protein secretion of Bacillus licheniformis compared to Bacillus subtilis.
J Mol Microbiol Biotechnol: 2009, 16(1-2);53-68
[PubMed:18957862]
[WorldCat.org]
[DOI]
(I p)
Yuping Wei, Teresa Havasy, Derrell C McPherson, David L Popham
Rod shape determination by the Bacillus subtilis class B penicillin-binding proteins encoded by pbpA and pbpH.
J Bacteriol: 2003, 185(16);4717-26
[PubMed:12896990]
[WorldCat.org]
[DOI]
(P p)
T Murray, D L Popham, C B Pearson, A R Hand, P Setlow
Analysis of outgrowth of Bacillus subtilis spores lacking penicillin-binding protein 2a.
J Bacteriol: 1998, 180(24);6493-502
[PubMed:9851991]
[WorldCat.org]
[DOI]
(P p)
T Murray, D L Popham, P Setlow
Identification and characterization of pbpA encoding Bacillus subtilis penicillin-binding protein 2A.
J Bacteriol: 1997, 179(9);3021-9
[PubMed:9139922]
[WorldCat.org]
[DOI]
(P p)