Difference between revisions of "OxdD"
| Line 20: | Line 20: | ||
|style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1176 bp, 392 aa | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 1176 bp, 392 aa | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yozS]]'', ''[[yoaO]]'' |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB13759]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB13759]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' | ||
Revision as of 11:35, 3 December 2012
- Description: oxalate decarboxylase, spore coat protein
| Gene name | oxdD |
| Synonyms | yoaN |
| Essential | no |
| Product | oxalate decarboxylase |
| Function | unknown |
| Gene expression levels in SubtiExpress: oxdD | |
| MW, pI | 43 kDa, 5.359 |
| Gene length, protein length | 1176 bp, 392 aa |
| Immediate neighbours | yozS, yoaO |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 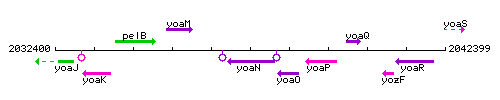
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed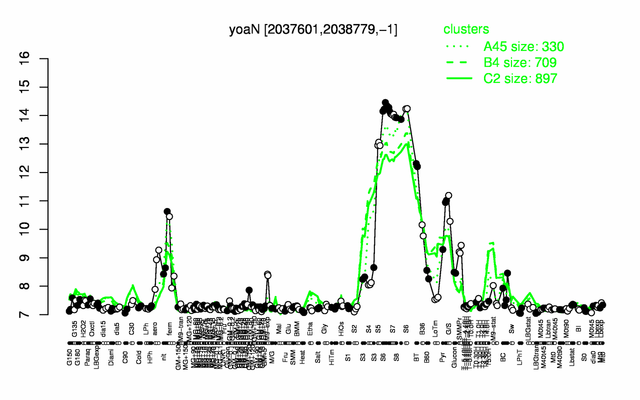
| |
Contents
Categories containing this gene/protein
sporulation proteins, resistance against other toxic compounds (nitric oxide, phenolic acids, flavonoids, oxalate)
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU18670
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Oxalate = formate + CO2 (according to Swiss-Prot)
- Protein family: UPF0361 family (according to Swiss-Prot)
- Paralogous protein(s): OxdC
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O34767
- KEGG entry: [3]
- E.C. number: 4.1.1.2
Additional information
Expression and regulation
- Operon: yoaN PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Miia R Mäkelä, Kristiina Hildén, Taina K Lundell
Oxalate decarboxylase: biotechnological update and prevalence of the enzyme in filamentous fungi.
Appl Microbiol Biotechnol: 2010, 87(3);801-14
[PubMed:20464388]
[WorldCat.org]
[DOI]
(I p)
Original publications
Sébastien Potot, Cláudia R Serra, Adriano O Henriques, Ghislain Schyns
Display of recombinant proteins on Bacillus subtilis spores, using a coat-associated enzyme as the carrier.
Appl Environ Microbiol: 2010, 76(17);5926-33
[PubMed:20601499]
[WorldCat.org]
[DOI]
(I p)
Leif Steil, Mónica Serrano, Adriano O Henriques, Uwe Völker
Genome-wide analysis of temporally regulated and compartment-specific gene expression in sporulating cells of Bacillus subtilis.
Microbiology (Reading): 2005, 151(Pt 2);399-420
[PubMed:15699190]
[WorldCat.org]
[DOI]
(P p)
Teresa Costa, Leif Steil, Lígia O Martins, Uwe Völker, Adriano O Henriques
Assembly of an oxalate decarboxylase produced under sigmaK control into the Bacillus subtilis spore coat.
J Bacteriol: 2004, 186(5);1462-74
[PubMed:14973022]
[WorldCat.org]
[DOI]
(P p)
A Tanner, L Bowater, S A Fairhurst, S Bornemann
Oxalate decarboxylase requires manganese and dioxygen for activity. Overexpression and characterization of Bacillus subtilis YvrK and YoaN.
J Biol Chem: 2001, 276(47);43627-34
[PubMed:11546787]
[WorldCat.org]
[DOI]
(P p)