Difference between revisions of "NsrR"
| Line 61: | Line 61: | ||
=== Additional information=== | === Additional information=== | ||
| − | |||
| − | |||
| − | |||
=The protein= | =The protein= | ||
| Line 112: | Line 109: | ||
* '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=nsrR_1013411_1013851_-1 nsrR] {{PubMed|22383849}} | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=nsrR_1013411_1013851_-1 nsrR] {{PubMed|22383849}} | ||
| − | * '''Sigma factor:''' | + | * '''[[Sigma factor]]:''' |
* '''Regulation:''' | * '''Regulation:''' | ||
| Line 145: | Line 142: | ||
<pubmed> 22287527 </pubmed> | <pubmed> 22287527 </pubmed> | ||
==Other original publications== | ==Other original publications== | ||
| − | <pubmed>16885456,,17293416,21091510 , </pubmed> | + | <pubmed>16885456, 24214949,17293416,21091510 , 19006327</pubmed> |
| − | |||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 10:57, 26 May 2014
- Description: nitric oxide-responsive regulator
| Gene name | nsrR |
| Synonyms | yhdE |
| Essential | no |
| Product | transcriptional repressor |
| Function | repression of ResD-ResE-dependent genes in the absence of nitric oxide (NO) |
| Gene expression levels in SubtiExpress: nsrR | |
| Metabolic function and regulation of this protein in SubtiPathways: nsrR | |
| MW, pI | 16 kDa, 7.75 |
| Gene length, protein length | 438 bp, 146 aa |
| Immediate neighbours | lytF, ygxB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 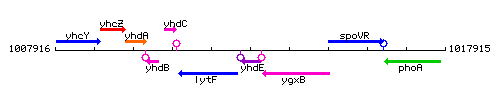
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed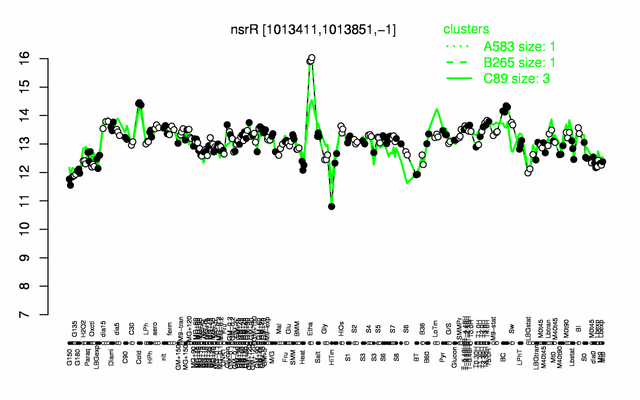
| |
Contents
Categories containing this gene/protein
transcription factors and their control, resistance against other toxic compounds (nitric oxide, phenolic acids, flavonoids, oxalate)
This gene is a member of the following regulons
The NsrR regulon
The gene
Basic information
- Locus tag: BSU09380
Phenotypes of a mutant
Database entries
- BsubCyc: BSU09380
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): [4Fe-4S] cluster, required for DNA-binding PubMed
- Effectors of protein activity:
Database entries
- BsubCyc: BSU09380
- Structure:
- UniProt: O07573
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Operon: nsrR PubMed
- Regulation:
- Regulatory mechanism:
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 122 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Elisabeth Härtig, Dieter Jahn
Regulation of the anaerobic metabolism in Bacillus subtilis.
Adv Microb Physiol: 2012, 61;195-216
[PubMed:23046954]
[WorldCat.org]
[DOI]
(I p)
The NsrR regulon
Sushma Kommineni, Amrita Lama, Benjamin Popescu, Michiko M Nakano
Global transcriptional control by NsrR in Bacillus subtilis.
J Bacteriol: 2012, 194(7);1679-88
[PubMed:22287527]
[WorldCat.org]
[DOI]
(I p)
Other original publications
Bernadette Henares, Sushma Kommineni, Onuma Chumsakul, Naotake Ogasawara, Shu Ishikawa, Michiko M Nakano
The ResD response regulator, through functional interaction with NsrR and fur, plays three distinct roles in Bacillus subtilis transcriptional control.
J Bacteriol: 2014, 196(2);493-503
[PubMed:24214949]
[WorldCat.org]
[DOI]
(I p)
Sushma Kommineni, Erik Yukl, Takahiro Hayashi, Jacob Delepine, Hao Geng, Pierre Moënne-Loccoz, Michiko M Nakano
Nitric oxide-sensitive and -insensitive interaction of Bacillus subtilis NsrR with a ResDE-controlled promoter.
Mol Microbiol: 2010, 78(5);1280-93
[PubMed:21091510]
[WorldCat.org]
[DOI]
(I p)
Erik T Yukl, Mohamed A Elbaz, Michiko M Nakano, Pierre Moënne-Loccoz
Transcription Factor NsrR from Bacillus subtilis Senses Nitric Oxide with a 4Fe-4S Cluster (†).
Biochemistry: 2008, 47(49);13084-92
[PubMed:19006327]
[WorldCat.org]
[DOI]
(I p)
Annika Rogstam, Jonas T Larsson, Peter Kjelgaard, Claes von Wachenfeldt
Mechanisms of adaptation to nitrosative stress in Bacillus subtilis.
J Bacteriol: 2007, 189(8);3063-71
[PubMed:17293416]
[WorldCat.org]
[DOI]
(P p)
Michiko M Nakano, Hao Geng, Shunji Nakano, Kazuo Kobayashi
The nitric oxide-responsive regulator NsrR controls ResDE-dependent gene expression.
J Bacteriol: 2006, 188(16);5878-87
[PubMed:16885456]
[WorldCat.org]
[DOI]
(P p)