NfrA
- Description: Spx-dependent FMN-containing NADPH-linked nitro/flavin reductase, stress protein
| Gene name | nfrA |
| Synonyms | ywcG, ipa-43d |
| Essential | no |
| Product | FMN-containing NADPH-linked nitro/flavin reductase |
| Function | unknown |
| Gene expression levels in SubtiExpress: nfrA | |
| Metabolic function and regulation of this protein in SubtiPathways: nfrA | |
| MW, pI | 28 kDa, 5.732 |
| Gene length, protein length | 747 bp, 249 aa |
| Immediate neighbours | ywcH, rodA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 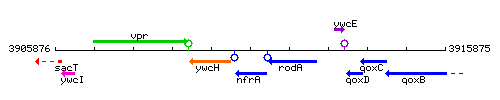
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed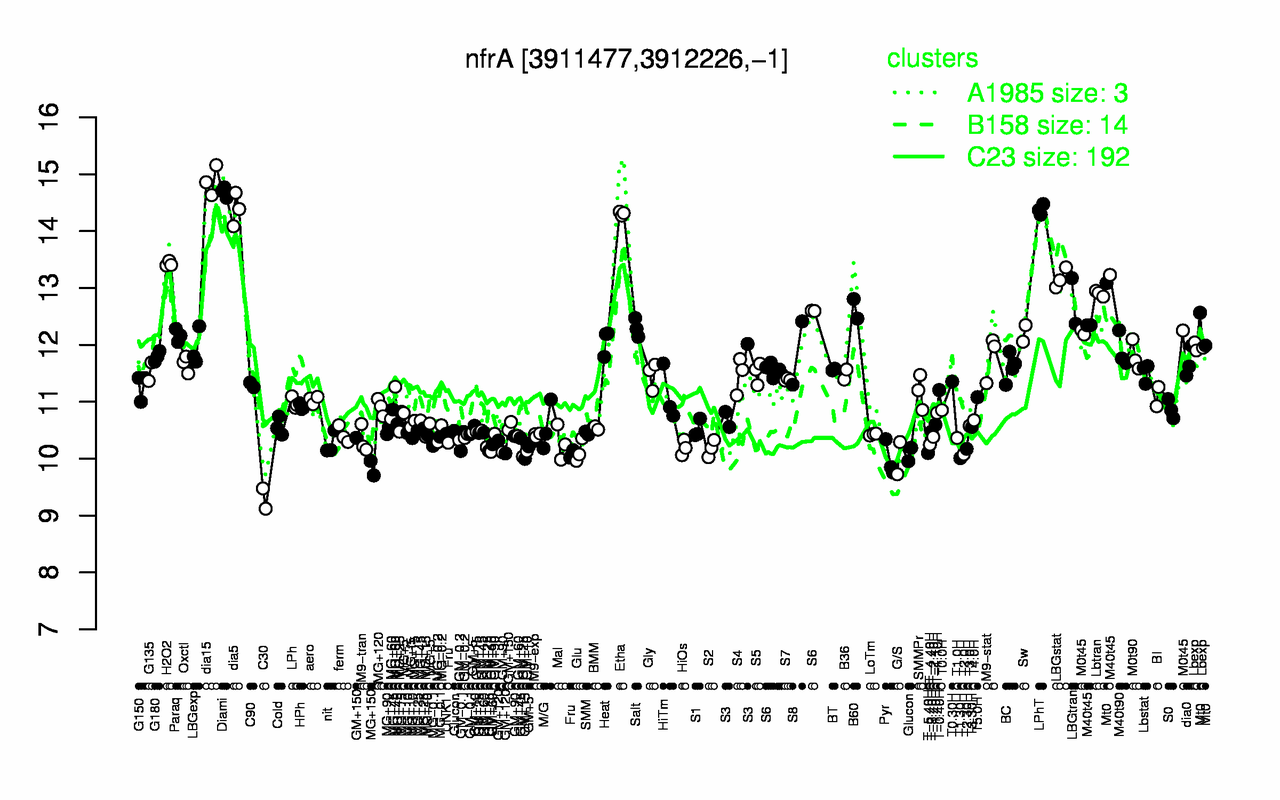
| |
Contents
Categories containing this gene/protein
electron transport/ other, resistance against oxidative and electrophile stress
This gene is a member of the following regulons
SigD regulon, Spo0A regulon, Spx regulon
The gene
Basic information
- Locus tag: BSU38110
Phenotypes of a mutant
Database entries
- BsubCyc: BSU38110
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: possesses an NADH oxidase activity that leads to high concentrations of oxygen peroxide and an NAD(+) degrading activity leading to free nicotinamide PubMed
- Protein family: flavin oxidoreductase frp family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU38110
- UniProt: P39605
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium): 222 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 616 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 560 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 960 PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Sylvie Cortial, Philippe Chaignon, Bogdan I Iorga, Stéphane Aymerich, Gilles Truan, Virginie Gueguen-Chaignon, Philippe Meyer, Solange Moréra, Jamal Ouazzani
NADH oxidase activity of Bacillus subtilis nitroreductase NfrA1: insight into its biological role.
FEBS Lett: 2010, 584(18);3916-22
[PubMed:20727352]
[WorldCat.org]
[DOI]
(I p)
Le Thi Tam, Haike Antelmann, Christine Eymann, Dirk Albrecht, Jörg Bernhardt, Michael Hecker
Proteome signatures for stress and starvation in Bacillus subtilis as revealed by a 2-D gel image color coding approach.
Proteomics: 2006, 6(16);4565-85
[PubMed:16847875]
[WorldCat.org]
[DOI]
(P p)
Virginie Molle, Masaya Fujita, Shane T Jensen, Patrick Eichenberger, José E González-Pastor, Jun S Liu, Richard Losick
The Spo0A regulon of Bacillus subtilis.
Mol Microbiol: 2003, 50(5);1683-701
[PubMed:14651647]
[WorldCat.org]
[DOI]
(P p)
Shunji Nakano, Michiko M Nakano, Ying Zhang, Montira Leelakriangsak, Peter Zuber
A regulatory protein that interferes with activator-stimulated transcription in bacteria.
Proc Natl Acad Sci U S A: 2003, 100(7);4233-8
[PubMed:12642660]
[WorldCat.org]
[DOI]
(P p)
C Moch, O Schrögel, R Allmansberger
Transcription of the nfrA-ywcH operon from Bacillus subtilis is specifically induced in response to heat.
J Bacteriol: 2000, 182(16);4384-93
[PubMed:10913069]
[WorldCat.org]
[DOI]
(P p)
S Zenno, T Kobori, M Tanokura, K Saigo
Purification and characterization of NfrA1, a Bacillus subtilis nitro/flavin reductase capable of interacting with the bacterial luciferase.
Biosci Biotechnol Biochem: 1998, 62(10);1978-87
[PubMed:9836433]
[WorldCat.org]
[DOI]
(P p)
C Moch, O Schrögel, R Allmansberger
The sigmaD-dependent transcription of the ywcG gene from Bacillus subtilis is dependent on an excess of glucose and glutamate.
Mol Microbiol: 1998, 27(5);889-98
[PubMed:9535080]
[WorldCat.org]
[DOI]
(P p)