Difference between revisions of "MurE"
| Line 62: | Line 62: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU15180&redirect=T BSU15180] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/murE-mraY-murD-spoVE-murG-murB-divIB-ylxWX-sbp.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/murE-mraY-murD-spoVE-murG-murB-divIB-ylxWX-sbp.html] | ||
| Line 100: | Line 101: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU15180&redirect=T BSU15180] | ||
* '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=2WTZ 2WTZ] (from ''Mycobacterium tuberculosis'', 35% identity, 50% similarity) {{PubMed|19945347}} | * '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=2WTZ 2WTZ] (from ''Mycobacterium tuberculosis'', 35% identity, 50% similarity) {{PubMed|19945347}} | ||
Revision as of 13:38, 2 April 2014
- Description: UDP-N-acetylmuramoyl-L-alanyl-D-glutamyl-meso-2,6-diaminopimelate synthetase
| Gene name | murE |
| Synonyms | |
| Essential | yes PubMed |
| Product | UDP-N-acetylmuramoyl-L-alanyl-D-glutamyl- meso-2,6-diaminopimelate synthetase |
| Function | peptidoglycan precursor biosynthesis |
| Gene expression levels in SubtiExpress: murE | |
| Metabolic function and regulation of this protein in SubtiPathways: murE | |
| MW, pI | 54 kDa, 5.558 |
| Gene length, protein length | 1482 bp, 494 aa |
| Immediate neighbours | spoVD, mraY |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 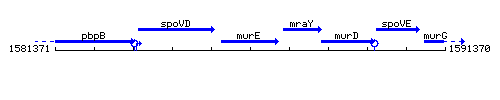
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed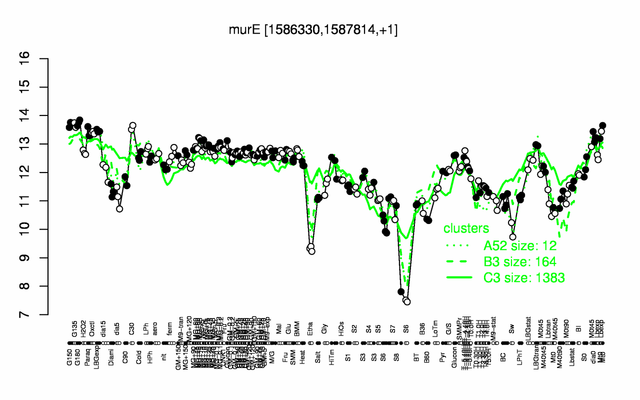
| |
Contents
Categories containing this gene/protein
cell wall synthesis, biosynthesis of cell wall components, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU15180
Phenotypes of a mutant
essential PubMed
Database entries
- BsubCyc: BSU15180
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + UDP-N-acetylmuramoyl-L-alanyl-D-glutamate + meso-2,6-diaminoheptanedioate = ADP + phosphate + UDP-N-acetylmuramoyl-L-alanyl-D-gamma-glutamyl-meso-2,6-diamino-heptanedioate (according to Swiss-Prot)
- Protein family: MurE subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm (according to Swiss-Prot)
Database entries
- BsubCyc: BSU15180
- UniProt: Q03523
- KEGG entry: [3]
- E.C. number: 6.3.2.13
Additional information
Expression and regulation
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Jean van Heijenoort
Lipid intermediates in the biosynthesis of bacterial peptidoglycan.
Microbiol Mol Biol Rev: 2007, 71(4);620-35
[PubMed:18063720]
[WorldCat.org]
[DOI]
(P p)
Christine Eymann, Georg Homuth, Christian Scharf, Michael Hecker
Bacillus subtilis functional genomics: global characterization of the stringent response by proteome and transcriptome analysis.
J Bacteriol: 2002, 184(9);2500-20
[PubMed:11948165]
[WorldCat.org]
[DOI]
(P p)
R A Daniel, J Errington
DNA sequence of the murE-murD region of Bacillus subtilis 168.
J Gen Microbiol: 1993, 139(2);361-70
[PubMed:8436954]
[WorldCat.org]
[DOI]
(P p)
A O Henriques, H de Lencastre, P J Piggot
A Bacillus subtilis morphogene cluster that includes spoVE is homologous to the mra region of Escherichia coli.
Biochimie: 1992, 74(7-8);735-48
[PubMed:1391053]
[WorldCat.org]
[DOI]
(P p)