Difference between revisions of "Mfd"
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || promotes strand-specific DNA repair by displacing | |style="background:#ABCDEF;" align="center"|'''Function''' || promotes strand-specific DNA repair by displacing | ||
| + | [[RNA polymerase]] stalled at a nucleotide lesion and directing | ||
| + | |||
| + | the (A)BC excinuclease to the RNA damage site | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://cellpublisher.gobics.de/subtiexpress/ ''Subti''Express]''': [http://cellpublisher.gobics.de/subtiexpress/bsu/BSU00550 mfd] | ||
[[RNA polymerase]] stalled at a nucleotide lesion and directing | [[RNA polymerase]] stalled at a nucleotide lesion and directing | ||
Revision as of 08:51, 8 August 2012
- Description: transcription-repair coupling factor
| Gene name | mfd |
| Synonyms | |
| Essential | no |
| Product | transcription-repair coupling factor |
| Function | promotes strand-specific DNA repair by displacing
RNA polymerase stalled at a nucleotide lesion and directing the (A)BC excinuclease to the RNA damage site |
| Gene expression levels in SubtiExpress: mfd
RNA polymerase stalled at a nucleotide lesion and directing the (A)BC excinuclease to the RNA damage site | |
| Interactions involving this protein in SubtInteract: Mfd | |
| MW, pI | 133 kDa, 5.367 |
| Gene length, protein length | 3531 bp, 1177 aa |
| Immediate neighbours | fin, spoVT |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 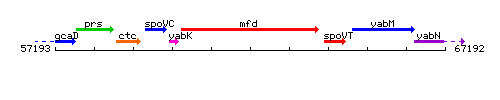
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, transcription
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU00550
Phenotypes of a mutant
In an mfd knock-out, the cell's ability to accumulate adaptive mutations in stationary phase is depressed. PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- promotes strand-specific DNA repair by displacing RNA polymerase stalled at a nucleotide lesion and directing the (A)BC excinuclease to the RNA damage site
- is required for roadblock transcription repression by transcription factors with binding sites downstream of the promoter (as for CcpA PubMed and CodY PubMed)
- Protein family:
- Paralogous protein(s): RecG
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P37474
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant: GP1167 (del ermC), available in Stülke lab
- Expression vector:
- lacZ fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Eduardo A Robleto, Holly A Martin, Mario Pedraza-Reyes
Mfd and transcriptional derepression cause genetic diversity in Bacillus subtilis.
Front Biosci (Elite Ed): 2012, 4(4);1246-54
[PubMed:22201950]
[WorldCat.org]
[DOI]
(I e)
Philip C Hanawalt, Graciela Spivak
Transcription-coupled DNA repair: two decades of progress and surprises.
Nat Rev Mol Cell Biol: 2008, 9(12);958-70
[PubMed:19023283]
[WorldCat.org]
[DOI]
(I p)
Rodrigo S Galhardo, P J Hastings, Susan M Rosenberg
Mutation as a stress response and the regulation of evolvability.
Crit Rev Biochem Mol Biol: 2007, 42(5);399-435
[PubMed:17917874]
[WorldCat.org]
[DOI]
(P p)
Alexandra M Deaconescu, Nigel Savery, Seth A Darst
The bacterial transcription repair coupling factor.
Curr Opin Struct Biol: 2007, 17(1);96-102
[PubMed:17239578]
[WorldCat.org]
[DOI]
(P p)
James J Truglio, Deborah L Croteau, Bennett Van Houten, Caroline Kisker
Prokaryotic nucleotide excision repair: the UvrABC system.
Chem Rev: 2006, 106(2);233-52
[PubMed:16464004]
[WorldCat.org]
[DOI]
(P p)
Sergei Borukhov, Jookyung Lee, Oleg Laptenko
Bacterial transcription elongation factors: new insights into molecular mechanism of action.
Mol Microbiol: 2005, 55(5);1315-24
[PubMed:15720542]
[WorldCat.org]
[DOI]
(P p)
Jeffrey Roberts, Joo-Seop Park
Mfd, the bacterial transcription repair coupling factor: translocation, repair and termination.
Curr Opin Microbiol: 2004, 7(2);120-5
[PubMed:15063847]
[WorldCat.org]
[DOI]
(P p)
E C Friedberg
Relationships between DNA repair and transcription.
Annu Rev Biochem: 1996, 65;15-42
[PubMed:8811173]
[WorldCat.org]
[DOI]
(P p)
C P Selby, A Sancar
Mechanisms of transcription-repair coupling and mutation frequency decline.
Microbiol Rev: 1994, 58(3);317-29
[PubMed:7968917]
[WorldCat.org]
[DOI]
(P p)
Original Articles
Additional publications: PubMed
Holly Anne Martin, Mario Pedraza-Reyes, Ronald E Yasbin, Eduardo A Robleto
Transcriptional de-repression and Mfd are mutagenic in stressed Bacillus subtilis cells.
J Mol Microbiol Biotechnol: 2011, 21(1-2);45-58
[PubMed:22248542]
[WorldCat.org]
[DOI]
(I p)
Katrin Gunka, Stefan Tholen, Jan Gerwig, Christina Herzberg, Jörg Stülke, Fabian M Commichau
A high-frequency mutation in Bacillus subtilis: requirements for the decryptification of the gudB glutamate dehydrogenase gene.
J Bacteriol: 2012, 194(5);1036-44
[PubMed:22178973]
[WorldCat.org]
[DOI]
(I p)
Boris R Belitsky, Abraham L Sonenshein
Roadblock repression of transcription by Bacillus subtilis CodY.
J Mol Biol: 2011, 411(4);729-43
[PubMed:21699902]
[WorldCat.org]
[DOI]
(I p)
Christine Pybus, Mario Pedraza-Reyes, Christian A Ross, Holly Martin, Katherine Ona, Ronald E Yasbin, Eduardo Robleto
Transcription-associated mutation in Bacillus subtilis cells under stress.
J Bacteriol: 2010, 192(13);3321-8
[PubMed:20435731]
[WorldCat.org]
[DOI]
(I p)
Christian Ross, Christine Pybus, Mario Pedraza-Reyes, Huang-Mo Sung, Ronald E Yasbin, Eduardo Robleto
Novel role of mfd: effects on stationary-phase mutagenesis in Bacillus subtilis.
J Bacteriol: 2006, 188(21);7512-20
[PubMed:16950921]
[WorldCat.org]
[DOI]
(P p)
Alexandra M Deaconescu, Anna L Chambers, Abigail J Smith, Bryce E Nickels, Ann Hochschild, Nigel J Savery, Seth A Darst
Structural basis for bacterial transcription-coupled DNA repair.
Cell: 2006, 124(3);507-20
[PubMed:16469698]
[WorldCat.org]
[DOI]
(P p)
J M Zalieckas, L V Wray, A E Ferson, S H Fisher
Transcription-repair coupling factor is involved in carbon catabolite repression of the Bacillus subtilis hut and gnt operons.
Mol Microbiol: 1998, 27(5);1031-8
[PubMed:9535092]
[WorldCat.org]
[DOI]
(P p)
S Ayora, F Rojo, N Ogasawara, S Nakai, J C Alonso
The Mfd protein of Bacillus subtilis 168 is involved in both transcription-coupled DNA repair and DNA recombination.
J Mol Biol: 1996, 256(2);301-18
[PubMed:8594198]
[WorldCat.org]
[DOI]
(P p)
V D Filippov, E E Zagoruiko
Study of MFD in Bacillus subtilis.
Mutat Res: 1978, 52(1);49-56
[PubMed:104170]
[WorldCat.org]
[DOI]
(P p)