Difference between revisions of "Mbl"
| Line 1: | Line 1: | ||
| − | * '''Description:''' cell- | + | * '''Description:''' [[cell shape]]-determining protein, forms filaments, the polymers control/restrict the mobility of the cell wall elongation enzyme complex <br/><br/> |
{| align="right" border="1" cellpadding="2" | {| align="right" border="1" cellpadding="2" | ||
| Line 10: | Line 10: | ||
|style="background:#ABCDEF;" align="center"| '''Essential''' || no | |style="background:#ABCDEF;" align="center"| '''Essential''' || no | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"| '''Product''' || MreB-like protein | + | |style="background:#ABCDEF;" align="center"| '''Product''' || [[MreB]]-like protein |
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Function''' || cell | + | |style="background:#ABCDEF;" align="center"|'''Function''' || [[cell shape]] determination |
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 35 kDa, 5.669 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 35 kDa, 5.669 | ||
| Line 81: | Line 81: | ||
* '''Effectors of protein activity:''' | * '''Effectors of protein activity:''' | ||
| − | * '''Interactions:''' [[MreB]]-[[Mbl]] | + | * '''[[SubtInteract|Interactions]]:''' |
| + | ** part of the [[cell wall biosynthetic complex]] {{PubMed|21636744,21636745}} | ||
| + | ** [[MreB]]-[[Mbl]] {{PubMed|21636744}} | ||
| + | ** [[Mbl]]-[[MreBH]] [http://www.ncbi.nlm.nih.gov/sites/entrez/17064365 PubMed] | ||
* '''Localization:''' | * '''Localization:''' | ||
| − | ** forms | + | ** during logarithmic growth, [[Mbl]] forms discrete patches thst move processively along peripheral tracks perpendicular to the cell axis {{PubMed|21636744}} |
| − | ** | + | ** forms transverse bands as cells enter the stationary phase {{PubMed|21636744}} |
| + | ** close to the inner surface of the cytoplasmic membrane [http://www.ncbi.nlm.nih.gov/sites/entrez/16950129 PubMed] | ||
| + | ** reports on helical structures formed by Mbl {{PubMed|16950129,20566861}} seem to be misinterpretation of data {{PubMed|21636744}} | ||
=== Database entries === | === Database entries === | ||
| Line 135: | Line 140: | ||
=References= | =References= | ||
==Localization== | ==Localization== | ||
| − | <pubmed> 17064365, 20566861 | + | <pubmed> 17064365, 20566861 21636744,21636745 </pubmed> |
==Other original publications== | ==Other original publications== | ||
<pubmed>19643765,16832063,11290328,12809607,16101995,12530960,7836311,,16950129 9023218, 19659933 19114499, 21091501 20525796</pubmed> | <pubmed>19643765,16832063,11290328,12809607,16101995,12530960,7836311,,16950129 9023218, 19659933 19114499, 21091501 20525796</pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 06:20, 11 July 2011
- Description: cell shape-determining protein, forms filaments, the polymers control/restrict the mobility of the cell wall elongation enzyme complex
| Gene name | mbl |
| Synonyms | |
| Essential | no |
| Product | MreB-like protein |
| Function | cell shape determination |
| MW, pI | 35 kDa, 5.669 |
| Gene length, protein length | 999 bp, 333 aa |
| Immediate neighbours | flhO, spoIIID |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 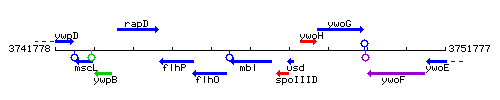
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
cell shape, sporulation proteins, membrane proteins
This gene is a member of the following regulons
SigE regulon, stringent response
The gene
Basic information
- Locus tag: BSU36410
Phenotypes of a mutant
non-viable in the presence of low Mg(2+), readily accumulate rsgI suppressor mutants PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- forms helical filaments in a heterologous system PubMed
- Protein family: ftsA/mreB family (according to Swiss-Prot)
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- during logarithmic growth, Mbl forms discrete patches thst move processively along peripheral tracks perpendicular to the cell axis PubMed
- forms transverse bands as cells enter the stationary phase PubMed
- close to the inner surface of the cytoplasmic membrane PubMed
- reports on helical structures formed by Mbl PubMed seem to be misinterpretation of data PubMed
Database entries
- Structure:
- UniProt: P39751
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
- An antisense RNA is predicted for mbl PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system: B. pertussis adenylate cyclase-based bacterial two hybrid system (BACTH), available in Görke lab
- Antibody: available in the Jeff Errington and Peter Graumann labs
Labs working on this gene/protein
Peter Graumann, Freiburg University, Germany homepage
Your additional remarks
References
Localization
Ethan C Garner, Remi Bernard, Wenqin Wang, Xiaowei Zhuang, David Z Rudner, Tim Mitchison
Coupled, circumferential motions of the cell wall synthesis machinery and MreB filaments in B. subtilis.
Science: 2011, 333(6039);222-5
[PubMed:21636745]
[WorldCat.org]
[DOI]
(I p)
Julia Domínguez-Escobar, Arnaud Chastanet, Alvaro H Crevenna, Vincent Fromion, Roland Wedlich-Söldner, Rut Carballido-López
Processive movement of MreB-associated cell wall biosynthetic complexes in bacteria.
Science: 2011, 333(6039);225-8
[PubMed:21636744]
[WorldCat.org]
[DOI]
(I p)
Henrik Strahl, Leendert W Hamoen
Membrane potential is important for bacterial cell division.
Proc Natl Acad Sci U S A: 2010, 107(27);12281-6
[PubMed:20566861]
[WorldCat.org]
[DOI]
(I p)
Hervé Joël Defeu Soufo, Peter L Graumann
Dynamic localization and interaction with other Bacillus subtilis actin-like proteins are important for the function of MreB.
Mol Microbiol: 2006, 62(5);1340-56
[PubMed:17064365]
[WorldCat.org]
[DOI]
(P p)
Other original publications
Hervé Joël Defeu Soufo, Peter L Graumann
Bacillus subtilis MreB paralogues have different filament architectures and lead to shape remodelling of a heterologous cell system.
Mol Microbiol: 2010, 78(5);1145-58
[PubMed:21091501]
[WorldCat.org]
[DOI]
(I p)
Irnov Irnov, Cynthia M Sharma, Jörg Vogel, Wade C Winkler
Identification of regulatory RNAs in Bacillus subtilis.
Nucleic Acids Res: 2010, 38(19);6637-51
[PubMed:20525796]
[WorldCat.org]
[DOI]
(I p)
Yoshikazu Kawai, Kei Asai, Jeffery Errington
Partial functional redundancy of MreB isoforms, MreB, Mbl and MreBH, in cell morphogenesis of Bacillus subtilis.
Mol Microbiol: 2009, 73(4);719-31
[PubMed:19659933]
[WorldCat.org]
[DOI]
(I p)
Kathrin Schirner, Jeff Errington
Influence of heterologous MreB proteins on cell morphology of Bacillus subtilis.
Microbiology (Reading): 2009, 155(Pt 11);3611-3621
[PubMed:19643765]
[WorldCat.org]
[DOI]
(I p)
Kathrin Schirner, Jeff Errington
The cell wall regulator {sigma}I specifically suppresses the lethal phenotype of mbl mutants in Bacillus subtilis.
J Bacteriol: 2009, 191(5);1404-13
[PubMed:19114499]
[WorldCat.org]
[DOI]
(I p)
Rut Carballido-López, Alex Formstone, Ying Li, S Dusko Ehrlich, Philippe Noirot, Jeff Errington
Actin homolog MreBH governs cell morphogenesis by localization of the cell wall hydrolase LytE.
Dev Cell: 2006, 11(3);399-409
[PubMed:16950129]
[WorldCat.org]
[DOI]
(P p)
Kittichoat Tiyanont, Thierry Doan, Michael B Lazarus, Xiao Fang, David Z Rudner, Suzanne Walker
Imaging peptidoglycan biosynthesis in Bacillus subtilis with fluorescent antibiotics.
Proc Natl Acad Sci U S A: 2006, 103(29);11033-8
[PubMed:16832063]
[WorldCat.org]
[DOI]
(P p)
Mark Leaver, Jeff Errington
Roles for MreC and MreD proteins in helical growth of the cylindrical cell wall in Bacillus subtilis.
Mol Microbiol: 2005, 57(5);1196-209
[PubMed:16101995]
[WorldCat.org]
[DOI]
(P p)
Richard A Daniel, Jeff Errington
Control of cell morphogenesis in bacteria: two distinct ways to make a rod-shaped cell.
Cell: 2003, 113(6);767-76
[PubMed:12809607]
[WorldCat.org]
[DOI]
(P p)
Rut Carballido-López, Jeff Errington
The bacterial cytoskeleton: in vivo dynamics of the actin-like protein Mbl of Bacillus subtilis.
Dev Cell: 2003, 4(1);19-28
[PubMed:12530960]
[WorldCat.org]
[DOI]
(P p)
L J Jones, R Carballido-López, J Errington
Control of cell shape in bacteria: helical, actin-like filaments in Bacillus subtilis.
Cell: 2001, 104(6);913-22
[PubMed:11290328]
[WorldCat.org]
[DOI]
(P p)
A Decatur, M T McMurry, B N Kunkel, R Losick
Translation of the mRNA for the sporulation gene spoIIID of Bacillus subtilis is dependent upon translation of a small upstream open reading frame.
J Bacteriol: 1997, 179(4);1324-8
[PubMed:9023218]
[WorldCat.org]
[DOI]
(P p)
Y Abhayawardhane, G C Stewart
Bacillus subtilis possesses a second determinant with extensive sequence similarity to the Escherichia coli mreB morphogene.
J Bacteriol: 1995, 177(3);765-73
[PubMed:7836311]
[WorldCat.org]
[DOI]
(P p)