Difference between revisions of "ManP"
(→References) |
|||
| Line 147: | Line 147: | ||
=References= | =References= | ||
| − | + | <pubmed>10627040 17218307 10627040, 20139185 23033921 23551403</pubmed> | |
| − | <pubmed>10627040 17218307 10627040, 20139185 </pubmed> | ||
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 20:20, 8 April 2013
- Description: trigger enzyme: mannose-specific phosphotransferase system, EIIBCA of the PTS
| Gene name | manP |
| Synonyms | yjdD |
| Essential | no |
| Product | trigger enzyme: mannose-specific phosphotransferase system, EIIBCA |
| Function | mannose uptake and phosphorylation, control of ManR activity |
| Gene expression levels in SubtiExpress: manP | |
| Interactions involving this protein in SubtInteract: ManP | |
| Metabolic function and regulation of this protein in SubtiPathways: Sugar catabolism | |
| MW, pI | 61 kDa, 6.715 |
| Gene length, protein length | 1767 bp, 589 aa |
| Immediate neighbours | manR, manA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 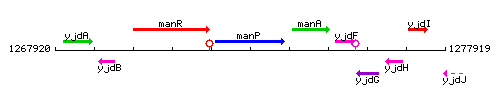
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed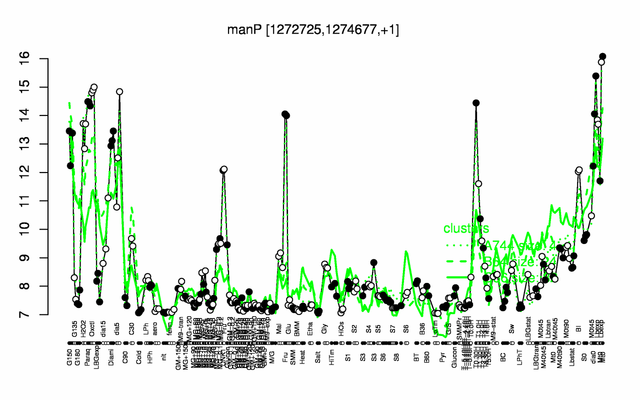
| |
Contents
Categories containing this gene/protein
phosphotransferase systems, utilization of specific carbon sources, transcription factors and their control, trigger enzyme, membrane proteins, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU12010
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: transport and concomitant phosphorylation of mannose
- Paralogous protein(s): FruA
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation on Ser-365 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization: membrane
Database entries
- Structure: 2R48 (IIB domain)
- UniProt: O31645
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Marian Wenzel, Josef Altenbuchner
The Bacillus subtilis mannose regulator, ManR, a DNA-binding protein regulated by HPr and its cognate PTS transporter ManP.
Mol Microbiol: 2013, 88(3);562-76
[PubMed:23551403]
[WorldCat.org]
[DOI]
(I p)
Imke G de Jong, Jan-Willem Veening, Oscar P Kuipers
Single cell analysis of gene expression patterns during carbon starvation in Bacillus subtilis reveals large phenotypic variation.
Environ Microbiol: 2012, 14(12);3110-21
[PubMed:23033921]
[WorldCat.org]
[DOI]
(I p)
Tianqi Sun, Josef Altenbuchner
Characterization of a mannose utilization system in Bacillus subtilis.
J Bacteriol: 2010, 192(8);2128-39
[PubMed:20139185]
[WorldCat.org]
[DOI]
(I p)
Boris Macek, Ivan Mijakovic, Jesper V Olsen, Florian Gnad, Chanchal Kumar, Peter R Jensen, Matthias Mann
The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis.
Mol Cell Proteomics: 2007, 6(4);697-707
[PubMed:17218307]
[WorldCat.org]
[DOI]
(P p)
Jonathan Reizer, Steffi Bachem, Aiala Reizer, Maryvonne Arnaud, Milton H Saier, Jörg Stülke
Novel phosphotransferase system genes revealed by genome analysis - the complete complement of PTS proteins encoded within the genome of Bacillus subtilis.
Microbiology (Reading): 1999, 145 ( Pt 12);3419-3429
[PubMed:10627040]
[WorldCat.org]
[DOI]
(P p)