Difference between revisions of "LutB"
(→Expression and regulation) |
(→This gene is a member of the following regulons) |
||
| Line 37: | Line 37: | ||
= This gene is a member of the following [[regulons]] = | = This gene is a member of the following [[regulons]] = | ||
| + | {{SubtiWiki regulon|[[FbpB regulon]]}}, | ||
{{SubtiWiki regulon|[[FsrA regulon]]}}, | {{SubtiWiki regulon|[[FsrA regulon]]}}, | ||
{{SubtiWiki regulon|[[LutR regulon]]}}, | {{SubtiWiki regulon|[[LutR regulon]]}}, | ||
Revision as of 15:27, 25 March 2012
- Description: lactate catabolic enzyme
| Gene name | lutB |
| Synonyms | yvfW |
| Essential | no |
| Product | lactate oxidase |
| Function | lactate utilization |
| Metabolic function and regulation of this protein in SubtiPathways: Central C-metabolism | |
| MW, pI | 53 kDa, 5.848 |
| Gene length, protein length | 1437 bp, 479 aa |
| Immediate neighbours | lutC, lutA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 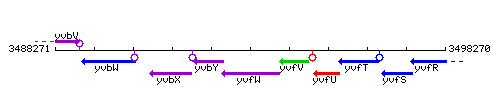
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
utilization of specific carbon sources
This gene is a member of the following regulons
FbpB regulon, FsrA regulon, LutR regulon, SinR regulon
The gene
Basic information
- Locus tag: BSU34040
Phenotypes of a mutant
no growth with lactate as the single carbon source PubMed
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: oxidation of lactate to pyruvate PubMed
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s): contains an iron-sulfur cluster
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O07021
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Richard Losick, Harvard Univ., Cambridge, USA homepage
Your additional remarks
References
Gregory T Smaldone, Haike Antelmann, Ahmed Gaballa, John D Helmann
The FsrA sRNA and FbpB protein mediate the iron-dependent induction of the Bacillus subtilis lutABC iron-sulfur-containing oxidases.
J Bacteriol: 2012, 194(10);2586-93
[PubMed:22427629]
[WorldCat.org]
[DOI]
(I p)
Gregory T Smaldone, Olga Revelles, Ahmed Gaballa, Uwe Sauer, Haike Antelmann, John D Helmann
A global investigation of the Bacillus subtilis iron-sparing response identifies major changes in metabolism.
J Bacteriol: 2012, 194(10);2594-605
[PubMed:22389480]
[WorldCat.org]
[DOI]
(I p)
Yunrong Chai, Roberto Kolter, Richard Losick
A widely conserved gene cluster required for lactate utilization in Bacillus subtilis and its involvement in biofilm formation.
J Bacteriol: 2009, 191(8);2423-30
[PubMed:19201793]
[WorldCat.org]
[DOI]
(I p)
Ahmed Gaballa, Haike Antelmann, Claudio Aguilar, Sukhjit K Khakh, Kyung-Bok Song, Gregory T Smaldone, John D Helmann
The Bacillus subtilis iron-sparing response is mediated by a Fur-regulated small RNA and three small, basic proteins.
Proc Natl Acad Sci U S A: 2008, 105(33);11927-32
[PubMed:18697947]
[WorldCat.org]
[DOI]
(I p)
Frances Chu, Daniel B Kearns, Steven S Branda, Roberto Kolter, Richard Losick
Targets of the master regulator of biofilm formation in Bacillus subtilis.
Mol Microbiol: 2006, 59(4);1216-28
[PubMed:16430695]
[WorldCat.org]
[DOI]
(P p)