Difference between revisions of "LiaR"
| Line 126: | Line 126: | ||
=References= | =References= | ||
| − | <pubmed>16816187, </pubmed> | + | <pubmed>15101989,17660417,16816187, </pubmed> |
# Jordan, S., A. Junker, J. D. Helmann, and T. Mascher. 2006. Regulation of LiaRS-dependent gene expression in Bacillus subtilis: identification of inhibitor proteins, regulator binding sites, and target genes of a conserved cell envelope stress-sensing two-component system. J. Bacteriol. 188: 5153-5166. [http://www.ncbi.nlm.nih.gov/sites/entrez/16816187 PubMed] | # Jordan, S., A. Junker, J. D. Helmann, and T. Mascher. 2006. Regulation of LiaRS-dependent gene expression in Bacillus subtilis: identification of inhibitor proteins, regulator binding sites, and target genes of a conserved cell envelope stress-sensing two-component system. J. Bacteriol. 188: 5153-5166. [http://www.ncbi.nlm.nih.gov/sites/entrez/16816187 PubMed] | ||
# Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] | ||
Revision as of 11:38, 12 June 2009
- Description: two-component response regulator, regulation of the liaI-liaH-liaG-liaF-liaS-liaR operon in response to bacitracin
| Gene name | liaR |
| Synonyms | yvqC |
| Essential | no |
| Product | two-component response regulator |
| Function | regulation of the liaI-liaH-liaG-liaF-liaS-liaR operon in response to bacitracin |
| MW, pI | 22 kDa, 4.956 |
| Gene length, protein length | 633 bp, 211 aa |
| Immediate neighbours | gerAC, liaS |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 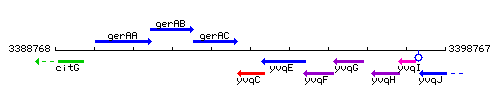
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU33080
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization: cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- Swiss prot entry: O32197
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Sigma factor: SigA
liaS: constitutive
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
John Helmann, Cornell University, USA Homepage
Thorsten Mascher, Karlsruhe Institute of Technology, Germany link
Your additional remarks
References
Sina Jordan, Eva Rietkötter, Mark A Strauch, Falk Kalamorz, Bronwyn G Butcher, John D Helmann, Thorsten Mascher
LiaRS-dependent gene expression is embedded in transition state regulation in Bacillus subtilis.
Microbiology (Reading): 2007, 153(Pt 8);2530-2540
[PubMed:17660417]
[WorldCat.org]
[DOI]
(P p)
Sina Jordan, Anja Junker, John D Helmann, Thorsten Mascher
Regulation of LiaRS-dependent gene expression in bacillus subtilis: identification of inhibitor proteins, regulator binding sites, and target genes of a conserved cell envelope stress-sensing two-component system.
J Bacteriol: 2006, 188(14);5153-66
[PubMed:16816187]
[WorldCat.org]
[DOI]
(P p)
Mélanie A Hamon, Nicola R Stanley, Robert A Britton, Alan D Grossman, Beth A Lazazzera
Identification of AbrB-regulated genes involved in biofilm formation by Bacillus subtilis.
Mol Microbiol: 2004, 52(3);847-60
[PubMed:15101989]
[WorldCat.org]
[DOI]
(P p)
- Jordan, S., A. Junker, J. D. Helmann, and T. Mascher. 2006. Regulation of LiaRS-dependent gene expression in Bacillus subtilis: identification of inhibitor proteins, regulator binding sites, and target genes of a conserved cell envelope stress-sensing two-component system. J. Bacteriol. 188: 5153-5166. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed