Difference between revisions of "LeuC"
| Line 60: | Line 60: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU28260&redirect=T BSU28260] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ilvBHC-leuABCD.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/ilvBHC-leuABCD.html] | ||
| Line 95: | Line 96: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU28260&redirect=T BSU28260] | ||
* '''Structure:''' | * '''Structure:''' | ||
Revision as of 14:25, 2 April 2014
- Description: 3-isopropylmalate dehydratase (large subunit)
| Gene name | leuC |
| Synonyms | |
| Essential | no |
| Product | 3-isopropylmalate dehydratase (large subunit) |
| Function | biosynthesis of leucine |
| Gene expression levels in SubtiExpress: leuC | |
| Metabolic function and regulation of this protein in SubtiPathways: leuC | |
| MW, pI | 52 kDa, 6.127 |
| Gene length, protein length | 1416 bp, 472 aa |
| Immediate neighbours | leuD, leuB |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 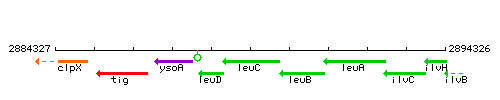
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed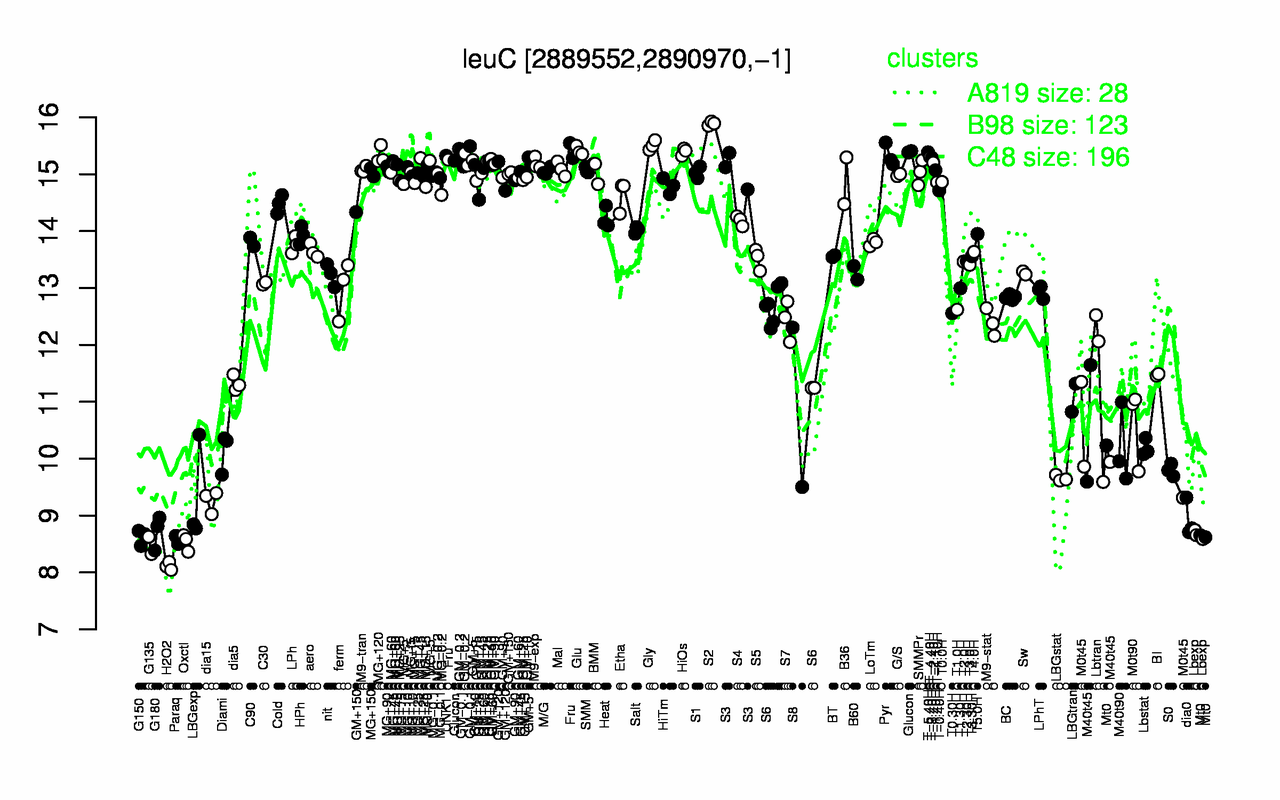
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids, phosphoproteins, most abundant proteins
This gene is a member of the following regulons
CcpA regulon, CodY regulon, FsrA regulon, T-box, TnrA regulon
The gene
Basic information
- Locus tag: BSU28260
Phenotypes of a mutant
Database entries
- BsubCyc: BSU28260
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (2R,3S)-3-isopropylmalate = (2S)-2-isopropylmaleate + H2O (according to Swiss-Prot)
- Protein family: LeuC type 1 subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- phosphorylated on Arg-81 PubMed
- Cofactors: contains an iron-sulfur cluster
- Effectors of protein activity:
Database entries
- BsubCyc: BSU28260
- Structure:
- UniProt: P80858
- KEGG entry: [3]
- E.C. number: 4.2.1.33
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Regulation:
- for a complete overview on the regulation of the ilv operon, see Brinsmade et al.
- repressed in the absence of good nitrogen sources (glutamine or ammonium) (TnrA) PubMed
- repressed during growth in the presence of branched chain amino acids (CodY) PubMed
- repressed by casamino acids PubMed
- expression is stimulated in the presence of glucose PubMed
- less expressed under conditions of extreme iron limitation (FsrA) PubMed
- Regulatory mechanism:
- Additional information:
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
- belongs to the 100 most abundant proteins PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References