Difference between revisions of "KhtS"
(→Expression and regulation) |
|||
| Line 99: | Line 99: | ||
=Expression and regulation= | =Expression and regulation= | ||
| − | * '''Operon:''' ''[[ | + | * '''Operon:''' ''[[khtS]]-[[khtT]]-[[khtU]]'' {{PubMed|22383849}} |
| − | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=khtS_1060988_1061326_-1 | + | * '''Expression browser:''' [http://genome.jouy.inra.fr/cgi-bin/seb/viewdetail.py?id=khtS_1060988_1061326_-1 khtS] {{PubMed|22383849}} |
* '''[[Sigma factor]]:''' | * '''[[Sigma factor]]:''' | ||
Revision as of 16:56, 14 February 2014
- Description: modulator of KhtU activity
| Gene name | yhaS |
| Synonyms | |
| Essential | no |
| Product | modulator of YhaU activity |
| Function | control of potassium ion efflux |
| Gene expression levels in SubtiExpress: khtS | |
| MW, pI | 12 kDa, 7.023 |
| Gene length, protein length | 336 bp, 112 aa |
| Immediate neighbours | khtT, yhaR |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 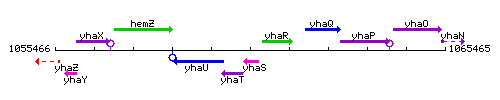
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed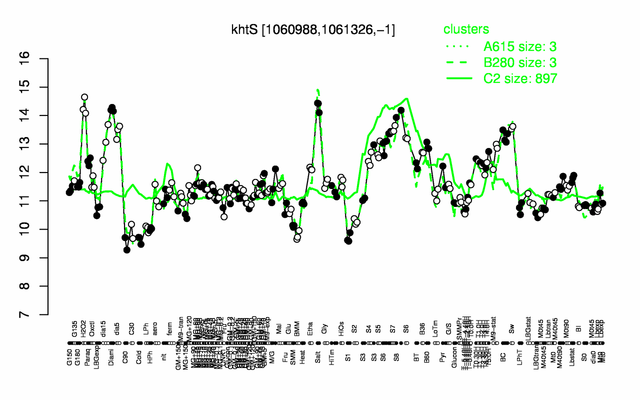
| |
Contents
Categories containing this gene/protein
metal ion homeostasis (K, Na, Ca, Mg)
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU09870
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Modification:
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: O07534
- KEGG entry: [2]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Pete Chandrangsu, Renata Dusi, Chris J Hamilton, John D Helmann
Methylglyoxal resistance in Bacillus subtilis: contributions of bacillithiol-dependent and independent pathways.
Mol Microbiol: 2014, 91(4);706-15
[PubMed:24330391]
[WorldCat.org]
[DOI]
(I p)
Makoto Fujisawa, Yuko Wada, Masahiro Ito
Modulation of the K+ efflux activity of Bacillus subtilis YhaU by YhaT and the C-terminal region of YhaS.
FEMS Microbiol Lett: 2004, 231(2);211-7
[PubMed:14987767]
[WorldCat.org]
[DOI]
(P p)