Difference between revisions of "HxlA"
m (Reverted edits by 134.76.70.252 (talk) to last revision by Umaeder) |
|||
| Line 13: | Line 13: | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"|'''Function''' || ribulose monophosphate pathway for formaldehyde fixation | |style="background:#ABCDEF;" align="center"|'''Function''' || ribulose monophosphate pathway for formaldehyde fixation | ||
| + | |- | ||
| + | |colspan="2" style="background:#FAF8CC;" align="center"| '''Gene expression levels in [http://cellpublisher.gobics.de/subtiexpress/ ''Subti''Express]''': [http://cellpublisher.gobics.de/subtiexpress/bsu/BSU03460 hxlA] | ||
|- | |- | ||
|style="background:#ABCDEF;" align="center"| '''MW, pI''' || 22 kDa, 4.564 | |style="background:#ABCDEF;" align="center"| '''MW, pI''' || 22 kDa, 4.564 | ||
Revision as of 09:39, 8 August 2012
- Description: 3-hexulose-6-phosphate synthase
| Gene name | hxlA |
| Synonyms | yckG |
| Essential | no |
| Product | 3-hexulose-6-phosphate synthase |
| Function | ribulose monophosphate pathway for formaldehyde fixation |
| Gene expression levels in SubtiExpress: hxlA | |
| MW, pI | 22 kDa, 4.564 |
| Gene length, protein length | 630 bp, 210 aa |
| Immediate neighbours | hxlB, hxlR |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 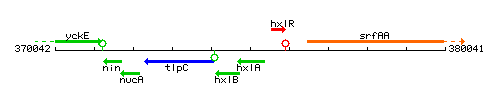
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed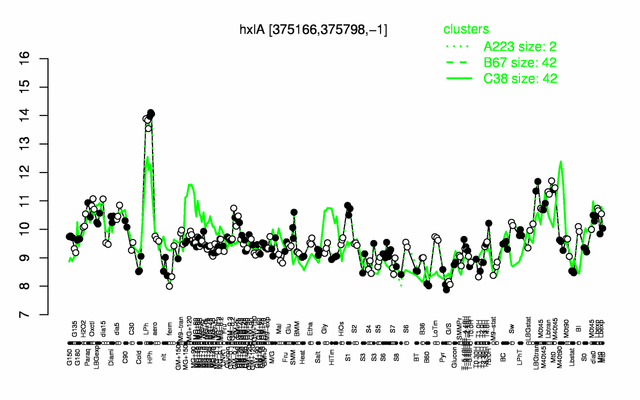
| |
Contents
Categories containing this gene/protein
resistance against oxidative and electrophile stress
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU03460
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: D-arabino-hex-3-ulose 6-phosphate = D-ribulose 5-phosphate + formaldehyde (according to Swiss-Prot)
- Protein family: HPS subfamily (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure:
- UniProt: P42405
- KEGG entry: [3]
- E.C. number: 4.1.2.43
Additional information
Expression and regulation
- Sigma factor:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Thi Thu Huyen Nguyen, Warawan Eiamphungporn, Ulrike Mäder, Manuel Liebeke, Michael Lalk, Michael Hecker, John D Helmann, Haike Antelmann
Genome-wide responses to carbonyl electrophiles in Bacillus subtilis: control of the thiol-dependent formaldehyde dehydrogenase AdhA and cysteine proteinase YraA by the MerR-family regulator YraB (AdhR).
Mol Microbiol: 2009, 71(4);876-94
[PubMed:19170879]
[WorldCat.org]
[DOI]
(I p)
Nobuo Kato, Hiroya Yurimoto, Rudolf K Thauer
The physiological role of the ribulose monophosphate pathway in bacteria and archaea.
Biosci Biotechnol Biochem: 2006, 70(1);10-21
[PubMed:16428816]
[WorldCat.org]
[DOI]
(P p)
H Yasueda, Y Kawahara, S Sugimoto
Bacillus subtilis yckG and yckF encode two key enzymes of the ribulose monophosphate pathway used by methylotrophs, and yckH is required for their expression.
J Bacteriol: 1999, 181(23);7154-60
[PubMed:10572115]
[WorldCat.org]
[DOI]
(P p)