Difference between revisions of "GyrB"
| Line 56: | Line 56: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU00060&redirect=T BSU00060] | ||
* '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yaaA-recF-yaaB-gyrB.html] | * '''DBTBS entry:''' [http://dbtbs.hgc.jp/COG/prom/yaaA-recF-yaaB-gyrB.html] | ||
| Line 95: | Line 96: | ||
=== Database entries === | === Database entries === | ||
| + | * '''BsubCyc:''' [http://bsubcyc.org/BSUB/NEW-IMAGE?type=NIL&object=BSU00060&redirect=T BSU00060] | ||
* '''Structure:''' | * '''Structure:''' | ||
Revision as of 12:44, 2 April 2014
- Description: DNA gyrase (subunit B)
| Gene name | gyrB |
| Synonyms | novA |
| Essential | yes PubMed |
| Product | DNA gyrase (subunit B) |
| Function | DNA supercoiling, initation of replication cycle and DNA elongation |
| Gene expression levels in SubtiExpress: GyrB | |
| Interactions involving this protein in SubtInteract: GyrB | |
| MW, pI | 71 kDa, 5.378 |
| Gene length, protein length | 1914 bp, 638 aa |
| Immediate neighbours | yaaB, gyrA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 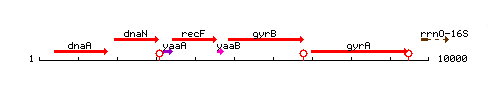
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed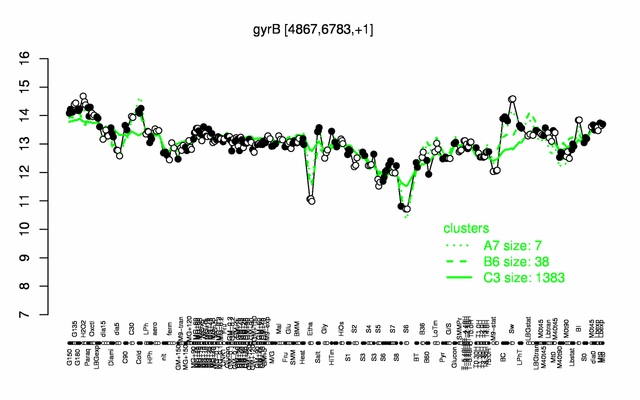
| |
Contents
Categories containing this gene/protein
DNA condensation/ segregation, essential genes
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU00060
Phenotypes of a mutant
essential PubMed
Database entries
- BsubCyc: BSU00060
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP-dependent breakage, passage and rejoining of double-stranded DNA (according to Swiss-Prot)
- Protein family: type II topoisomerase family (according to Swiss-Prot)
- Paralogous protein(s): ParE
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylated on Ser-400 PubMed
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- nucleoid-associated in actively growing cells PubMed
Database entries
- BsubCyc: BSU00060
- Structure:
- UniProt: P05652
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information: GyrB is subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Dagmar Klostermeier, Biozentrum Basel, Switzerland homepage
Your additional remarks
References
Additional references: PubMed
Luise A K Kleine Borgmann, Hanna Hummel, Maximilian H Ulbrich, Peter L Graumann
SMC condensation centers in Bacillus subtilis are dynamic structures.
J Bacteriol: 2013, 195(10);2136-45
[PubMed:23475963]
[WorldCat.org]
[DOI]
(I p)
Boumediene Soufi, Chanchal Kumar, Florian Gnad, Matthias Mann, Ivan Mijakovic, Boris Macek
Stable isotope labeling by amino acids in cell culture (SILAC) applied to quantitative proteomics of Bacillus subtilis.
J Proteome Res: 2010, 9(7);3638-46
[PubMed:20509597]
[WorldCat.org]
[DOI]
(I p)
Airat Gubaev, Manuel Hilbert, Dagmar Klostermeier
The DNA-gate of Bacillus subtilis gyrase is predominantly in the closed conformation during the DNA supercoiling reaction.
Proc Natl Acad Sci U S A: 2009, 106(32);13278-83
[PubMed:19666507]
[WorldCat.org]
[DOI]
(I p)
Ulf Gerth, Holger Kock, Ilja Kusters, Stephan Michalik, Robert L Switzer, Michael Hecker
Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis.
J Bacteriol: 2008, 190(1);321-31
[PubMed:17981983]
[WorldCat.org]
[DOI]
(I p)
Thomas Göttler, Dagmar Klostermeier
Dissection of the nucleotide cycle of B. subtilis DNA gyrase and its modulation by DNA.
J Mol Biol: 2007, 367(5);1392-404
[PubMed:17320901]
[WorldCat.org]
[DOI]
(P p)
W M Huang, J L Libbey, P van der Hoeven, S X Yu
Bipolar localization of Bacillus subtilis topoisomerase IV, an enzyme required for chromosome segregation.
Proc Natl Acad Sci U S A: 1998, 95(8);4652-7
[PubMed:9539793]
[WorldCat.org]
[DOI]
(P p)
N Ogasawara, S Moriya, H Yoshikawa
Structure and function of the region of the replication origin of the Bacillus subtilis chromosome. IV. Transcription of the oriC region and expression of DNA gyrase genes and other open reading frames.
Nucleic Acids Res: 1985, 13(7);2267-79
[PubMed:2987848]
[WorldCat.org]
[DOI]
(P p)