FadB
- Description: 3-hydroxyacyl-CoA dehydratase
| Gene name | fadB |
| Synonyms | ysiB |
| Essential | no |
| Product | 3-hydroxyacyl-CoA dehydratase |
| Function | fatty acid degradation |
| Gene expression levels in SubtiExpress: fadB | |
| Metabolic function and regulation of this protein in SubtiPathways: Fatty acid degradation | |
| MW, pI | 27 kDa, 4.951 |
| Gene length, protein length | 774 bp, 258 aa |
| Immediate neighbours | etfB, fadR |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 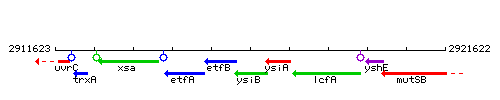
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed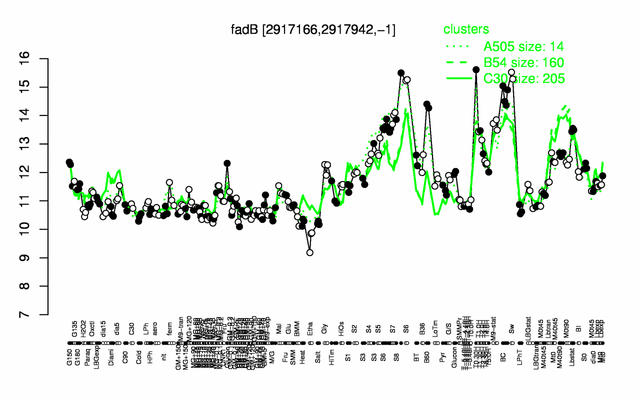
| |
Contents
Categories containing this gene/protein
utilization of lipids, phosphoproteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU28540
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: (3S)-3-hydroxyacyl-CoA = trans-2(or 3)-enoyl-CoA + H2O (according to Swiss-Prot)
- Protein family: enoyl-CoA hydratase/isomerase family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- phosphorylated on Arg-230 PubMed
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 3PEA (from B. anthracis, 49% identity, 66% similarity)
- UniProt: P94549
- KEGG entry: [3]
- E.C. number: 4.2.1.17
Additional information
Expression and regulation
- Regulation:
- induced by long-chain fatty acids (FadR) PubMed
- subject to carbon catabolite repression (3.8-fold) (CcpA-HPr(Ser-P) PubMed
- Regulatory mechanism:
- FadR: transcription repression PubMed
- CcpA-HPr(Ser-P): transcription repression PubMed
- Additional information:
Biological materials
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Alexander K W Elsholz, Kürsad Turgay, Stephan Michalik, Bernd Hessling, Katrin Gronau, Dan Oertel, Ulrike Mäder, Jörg Bernhardt, Dörte Becher, Michael Hecker, Ulf Gerth
Global impact of protein arginine phosphorylation on the physiology of Bacillus subtilis.
Proc Natl Acad Sci U S A: 2012, 109(19);7451-6
[PubMed:22517742]
[WorldCat.org]
[DOI]
(I p)
Yasutaro Fujita, Hiroshi Matsuoka, Kazutake Hirooka
Regulation of fatty acid metabolism in bacteria.
Mol Microbiol: 2007, 66(4);829-39
[PubMed:17919287]
[WorldCat.org]
[DOI]
(P p)
Hiroshi Matsuoka, Kazutake Hirooka, Yasutaro Fujita
Organization and function of the YsiA regulon of Bacillus subtilis involved in fatty acid degradation.
J Biol Chem: 2007, 282(8);5180-94
[PubMed:17189250]
[WorldCat.org]
[DOI]
(P p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)