Difference between revisions of "ExoA"
(→References) |
|||
| Line 35: | Line 35: | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
<br/><br/><br/><br/> | <br/><br/><br/><br/> | ||
| − | |||
| − | |||
| − | |||
| − | |||
<br/><br/> | <br/><br/> | ||
| Line 121: | Line 117: | ||
* '''Mutant:''' | * '''Mutant:''' | ||
| − | + | ** GP898 (''exoA''::''kan''), available in [[Jörg Stülke]]'s lab | |
| − | ** GP898 (''exoA''::''kan''), available in [[Stülke]] lab | + | ** GP1503 (''exoA''::''kan'', ''nfo''::''cat''), available in [[Jörg Stülke]]'s lab |
| − | ** GP1503 (''exoA''::''kan'', ''nfo''::''cat''), available in [[Stülke]] lab | ||
* '''Expression vector:''' | * '''Expression vector:''' | ||
| Line 143: | Line 138: | ||
<pubmed> 22933559 </pubmed> | <pubmed> 22933559 </pubmed> | ||
== Original publications == | == Original publications == | ||
| − | <pubmed>18203828,16237020, 19930460, 10540738 21441501</pubmed> | + | <pubmed>18203828,16237020, 19930460, 10540738 21441501 24123749 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 14:25, 21 October 2013
- Description: apurinic/apyrimidinic endonuclease, multifunctional DNA-repair enzyme, important for spore dormance
| Gene name | exoA |
| Synonyms | |
| Essential | no |
| Product | apurinic/apyrimidinic endonuclease |
| Function | repair of oxidative DNA damage in spores |
| Gene expression levels in SubtiExpress: exoA | |
| MW, pI | 29 kDa, 4.592 |
| Gene length, protein length | 756 bp, 252 aa |
| Immediate neighbours | ccpB, rpsR |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 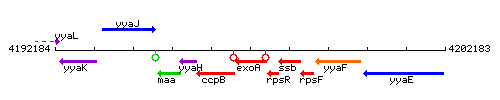
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed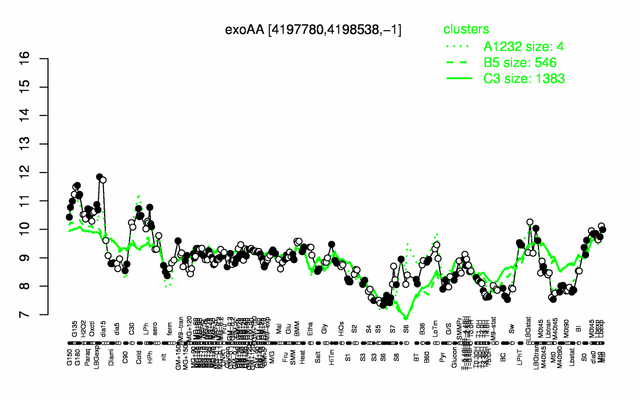
| |
Contents
Categories containing this gene/protein
DNA repair/ recombination, sporulation proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU40880
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Exonucleolytic cleavage in the 3'- to 5'-direction to yield nucleoside 5'-phosphates (according to Swiss-Prot)
- Protein family: DNA repair enzymes AP/exoA family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
- cytoplasm (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: P37454
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- GP898 (exoA::kan), available in Jörg Stülke's lab
- GP1503 (exoA::kan, nfo::cat), available in Jörg Stülke's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
Ralf Moeller, Marina Raguse, Günther Reitz, Ryuichi Okayasu, Zuofeng Li, Stuart Klein, Peter Setlow, Wayne L Nicholson
Resistance of Bacillus subtilis spore DNA to lethal ionizing radiation damage relies primarily on spore core components and DNA repair, with minor effects of oxygen radical detoxification.
Appl Environ Microbiol: 2014, 80(1);104-9
[PubMed:24123749]
[WorldCat.org]
[DOI]
(I p)
Ralf Moeller, Peter Setlow, Mario Pedraza-Reyes, Ryuichi Okayasu, Günther Reitz, Wayne L Nicholson
Role of the Nfo and ExoA apurinic/apyrimidinic endonucleases in radiation resistance and radiation-induced mutagenesis of Bacillus subtilis spores.
J Bacteriol: 2011, 193(11);2875-9
[PubMed:21441501]
[WorldCat.org]
[DOI]
(I p)
Marcelo Barraza-Salas, Juan R Ibarra-Rodríguez, Silvia J Mellado, José M Salas-Pacheco, Peter Setlow, Mario Pedraza-Reyes
Effects of forespore-specific overexpression of apurinic/apyrimidinic endonuclease Nfo on the DNA-damage resistance properties of Bacillus subtilis spores.
FEMS Microbiol Lett: 2010, 302(2);159-65
[PubMed:19930460]
[WorldCat.org]
[DOI]
(I p)
Juan R Ibarra, Alma D Orozco, Juan A Rojas, Karina López, Peter Setlow, Ronald E Yasbin, Mario Pedraza-Reyes
Role of the Nfo and ExoA apurinic/apyrimidinic endonucleases in repair of DNA damage during outgrowth of Bacillus subtilis spores.
J Bacteriol: 2008, 190(6);2031-8
[PubMed:18203828]
[WorldCat.org]
[DOI]
(I p)
José M Salas-Pacheco, Barbara Setlow, Peter Setlow, Mario Pedraza-Reyes
Role of the Nfo (YqfS) and ExoA apurinic/apyrimidinic endonucleases in protecting Bacillus subtilis spores from DNA damage.
J Bacteriol: 2005, 187(21);7374-81
[PubMed:16237020]
[WorldCat.org]
[DOI]
(P p)
T Shida, T Ogawa, N Ogasawara, J Sekiguchi
Characterization of Bacillus subtilis ExoA protein: a multifunctional DNA-repair enzyme similar to the Escherichia coli exonuclease III.
Biosci Biotechnol Biochem: 1999, 63(9);1528-34
[PubMed:10540738]
[WorldCat.org]
[DOI]
(P p)