Difference between revisions of "EpsJ"
(→Expression and regulation) |
|||
| Line 87: | Line 87: | ||
=Expression and regulation= | =Expression and regulation= | ||
| + | * '''Operon:''' ''[[epsA]]-[[epsB]]-[[epsC]]-[[epsD]]-[[epsE]]-[[epsF]]-[[epsG]]-[[epsH]]-[[epsI]]-[[epsJ]]-[[epsK]]-[[epsL]]-[[epsM]]-[[epsN]]-[[epsO]]'' {{PubMed|15661000}} | ||
| − | + | * '''[[Sigma factor]]:''' [[SigA]] {{PubMed|15661000}} | |
| − | |||
| − | * '''[[Sigma factor]]:''' [[SigA]] | ||
* '''Regulation:''' | * '''Regulation:''' | ||
| − | ** repressed by [[SinR]] | + | ** repressed by [[SinR]] {{PubMed|15661000}} |
| − | * '''Regulatory mechanism:''' [[SinR]]: transcription repression | + | * '''Regulatory mechanism:''' |
| − | ** [[ | + | ** [[SinR]]: transcription repression {{PubMed|15661000}} |
| + | ** [[SlrR]]: transcription activation {{PubMed|18647168}} | ||
| − | * '''Additional information:''' induction by sequestration of [[SinR]] by [[SinI]] [ | + | * '''Additional information:''' |
| + | ** induction by sequestration of [[SinR]] by [[SinI]] {{PubMed|15661000}} | ||
| + | ** activation by [[SlrR]] requires prior formation of the [[SlrR]]-[[SlrA]] complex {{PubMed|18647168}} | ||
=Biological materials = | =Biological materials = | ||
Revision as of 09:21, 30 December 2009
- Description: similar to 1,4-galactosyltransferase
| Gene name | epsJ |
| Synonyms | yveT |
| Essential | no |
| Product | unknown |
| Function | extracellular polysaccharide synthesis |
| MW, pI | 39 kDa, 7.567 |
| Gene length, protein length | 1032 bp, 344 aa |
| Immediate neighbours | epsK, epsI |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 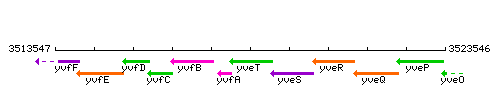
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU34280
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: glycosyltransferase 2 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- UniProt: P71059
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Richard Losick, Harvard Univ., Cambridge, USA homepage
Your additional remarks
References
Yunrong Chai, Frances Chu, Roberto Kolter, Richard Losick
Bistability and biofilm formation in Bacillus subtilis.
Mol Microbiol: 2008, 67(2);254-63
[PubMed:18047568]
[WorldCat.org]
[DOI]
(P p)
Frances Chu, Daniel B Kearns, Steven S Branda, Roberto Kolter, Richard Losick
Targets of the master regulator of biofilm formation in Bacillus subtilis.
Mol Microbiol: 2006, 59(4);1216-28
[PubMed:16430695]
[WorldCat.org]
[DOI]
(P p)
Daniel B Kearns, Frances Chu, Steven S Branda, Roberto Kolter, Richard Losick
A master regulator for biofilm formation by Bacillus subtilis.
Mol Microbiol: 2005, 55(3);739-49
[PubMed:15661000]
[WorldCat.org]
[DOI]
(P p)