Difference between revisions of "DivIB"
| Line 26: | Line 26: | ||
|style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[murB]]'', ''[[ylxW]]'' | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[murB]]'', ''[[ylxW]]'' | ||
|- | |- | ||
| − | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU15240 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU15240 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU15240 | + | |style="background:#FAF8CC;" align="center"|'''Sequences'''||[http://bsubcyc.org/BSUB/sequence-aa?type=GENE&object=BSU15240 Protein] [http://bsubcyc.org/BSUB/sequence?type=GENE&object=BSU15240 DNA] [http://bsubcyc.org/BSUB/seq-selector?chromosome=CHROM-1&object=BSU15240 DNA_with_flanks] |
|- | |- | ||
|colspan="2" | '''Genetic context''' <br/> [[Image:divIB_context.gif]] | |colspan="2" | '''Genetic context''' <br/> [[Image:divIB_context.gif]] | ||
Revision as of 10:09, 14 May 2013
- Description: cell-division initiation protein (septum formation)
| Gene name | divIB |
| Synonyms | dds, tms-12 |
| Essential | yes PubMed |
| Product | cell-division protein |
| Function | septum formation |
| Gene expression levels in SubtiExpress: divIB | |
| Interactions involving this protein in SubtInteract: DivIB | |
| Metabolic function and regulation of this protein in SubtiPathways: Cell wall | |
| MW, pI | 30 kDa, 9.244 |
| Gene length, protein length | 789 bp, 263 aa |
| Immediate neighbours | murB, ylxW |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 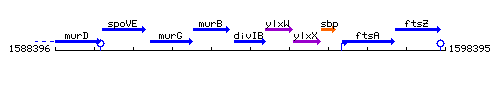
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed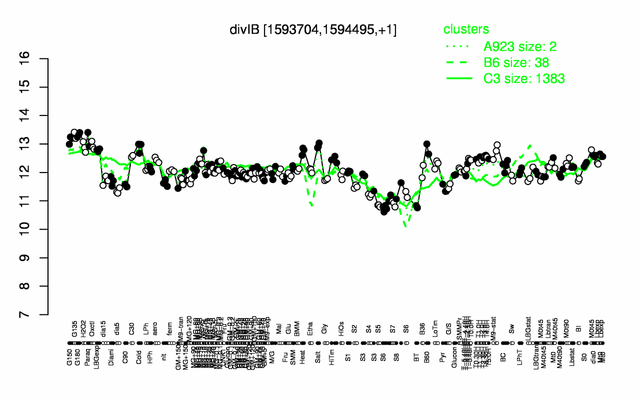
| |
Contents
Categories containing this gene/protein
cell division, sporulation proteins, cell envelope stress proteins (controlled by SigM, V, W, X, Y), essential genes, membrane proteins
This gene is a member of the following regulons
SigE regulon, SigM regulon, SpoIIID regulon
The gene
Basic information
- Locus tag: BSU15240
Phenotypes of a mutant
essential PubMed
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: ftsQ family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane PubMed
Database entries
- Structure: 2ALJ (major extracytoplasmic domain, Geobacillus stearothermophilus)
- UniProt: P16655
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional references: PubMed
Tatsuya Fukushima, Isako Furihata, Robyn Emmins, Richard A Daniel, James A Hoch, Hendrik Szurmant
A role for the essential YycG sensor histidine kinase in sensing cell division.
Mol Microbiol: 2011, 79(2);503-22
[PubMed:21219466]
[WorldCat.org]
[DOI]
(I p)
Pamela Gamba, Jan-Willem Veening, Nigel J Saunders, Leendert W Hamoen, Richard A Daniel
Two-step assembly dynamics of the Bacillus subtilis divisome.
J Bacteriol: 2009, 191(13);4186-94
[PubMed:19429628]
[WorldCat.org]
[DOI]
(I p)
Warawan Eiamphungporn, John D Helmann
The Bacillus subtilis sigma(M) regulon and its contribution to cell envelope stress responses.
Mol Microbiol: 2008, 67(4);830-48
[PubMed:18179421]
[WorldCat.org]
[DOI]
(P p)
L S Thompson, P L Beech, G Real, A O Henriques, E J Harry
Requirement for the cell division protein DivIB in polar cell division and engulfment during sporulation in Bacillus subtilis.
J Bacteriol: 2006, 188(21);7677-85
[PubMed:16936026]
[WorldCat.org]
[DOI]
(P p)
Richard A Daniel, Marie-Françoise Noirot-Gros, Philippe Noirot, Jeff Errington
Multiple interactions between the transmembrane division proteins of Bacillus subtilis and the role of FtsL instability in divisome assembly.
J Bacteriol: 2006, 188(21);7396-404
[PubMed:16936019]
[WorldCat.org]
[DOI]
(P p)
Gonçalo Real, Sabine Autret, Elizabeth J Harry, Jeffery Errington, Adriano O Henriques
Cell division protein DivIB influences the Spo0J/Soj system of chromosome segregation in Bacillus subtilis.
Mol Microbiol: 2005, 55(2);349-67
[PubMed:15659156]
[WorldCat.org]
[DOI]
(P p)
V L Katis, R G Wake, E J Harry
Septal localization of the membrane-bound division proteins of Bacillus subtilis DivIB and DivIC is codependent only at high temperatures and requires FtsZ.
J Bacteriol: 2000, 182(12);3607-11
[PubMed:10852898]
[WorldCat.org]
[DOI]
(P p)
R A Daniel, J Errington
Intrinsic instability of the essential cell division protein FtsL of Bacillus subtilis and a role for DivIB protein in FtsL turnover.
Mol Microbiol: 2000, 36(2);278-89
[PubMed:10792716]
[WorldCat.org]
[DOI]
(P p)
V L Katis, R G Wake
Membrane-bound division proteins DivIB and DivIC of Bacillus subtilis function solely through their external domains in both vegetative and sporulation division.
J Bacteriol: 1999, 181(9);2710-8
[PubMed:10217758]
[WorldCat.org]
[DOI]
(P p)
R A Daniel, E J Harry, V L Katis, R G Wake, J Errington
Characterization of the essential cell division gene ftsL(yIID) of Bacillus subtilis and its role in the assembly of the division apparatus.
Mol Microbiol: 1998, 29(2);593-604
[PubMed:9720875]
[WorldCat.org]
[DOI]
(P p)
V L Katis, E J Harry, R G Wake
The Bacillus subtilis division protein DivIC is a highly abundant membrane-bound protein that localizes to the division site.
Mol Microbiol: 1997, 26(5);1047-55
[PubMed:9426141]
[WorldCat.org]
[DOI]
(P p)
S L Rowland, V L Katis, S R Partridge, R G Wake
DivIB, FtsZ and cell division in Bacillus subtilis.
Mol Microbiol: 1997, 23(2);295-302
[PubMed:9044263]
[WorldCat.org]
[DOI]
(P p)
E J Harry, S L Rowland, M S Malo, R G Wake
Expression of divIB of Bacillus subtilis during vegetative growth.
J Bacteriol: 1994, 176(4);1172-9
[PubMed:8106328]
[WorldCat.org]
[DOI]
(P p)
G Theeragool, A Miyao, K Yamada, T Sato, Y Kobayashi
In vivo expression of the Bacillus subtilis spoVE gene.
J Bacteriol: 1993, 175(13);4071-80
[PubMed:8320223]
[WorldCat.org]
[DOI]
(P p)
E J Harry, B J Stewart, R G Wake
Characterization of mutations in divIB of Bacillus subtilis and cellular localization of the DivIB protein.
Mol Microbiol: 1993, 7(4);611-21
[PubMed:8459777]
[WorldCat.org]
[DOI]
(P p)
A O Henriques, H de Lencastre, P J Piggot
A Bacillus subtilis morphogene cluster that includes spoVE is homologous to the mra region of Escherichia coli.
Biochimie: 1992, 74(7-8);735-48
[PubMed:1391053]
[WorldCat.org]
[DOI]
(P p)