Difference between revisions of "DeoD"
| Line 22: | Line 22: | ||
|style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 699 bp, 233 aa | |style="background:#ABCDEF;" align="center"| '''Gene length, protein length''' || 699 bp, 233 aa | ||
|- | |- | ||
| − | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yodJ]]'', ''[[ | + | |style="background:#ABCDEF;" align="center"|'''Immediate neighbours''' || ''[[yodJ]]'', ''[[yoyE]]'' |
|- | |- | ||
|colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB13854]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' | |colspan="2" style="background:#FAF8CC;" align="center"|'''Get the DNA and protein [http://srs.ebi.ac.uk/srsbin/cgi-bin/wgetz?-e+[EMBLCDS:CAB13854]+-newId sequences] <br/> (Barbe ''et al.'', 2009)''' | ||
Revision as of 12:12, 3 December 2012
- Description: purine nucleoside phosphorylase
| Gene name | deoD |
| Synonyms | |
| Essential | no |
| Product | purine nucleoside phosphorylase |
| Function | purine salvage and interconversion |
| Gene expression levels in SubtiExpress: deoD | |
| Metabolic function and regulation of this protein in SubtiPathways: Purine salvage, Nucleoside catabolism | |
| MW, pI | 25 kDa, 4.978 |
| Gene length, protein length | 699 bp, 233 aa |
| Immediate neighbours | yodJ, yoyE |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 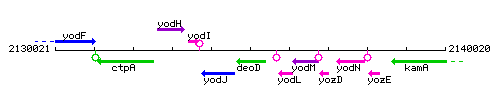
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed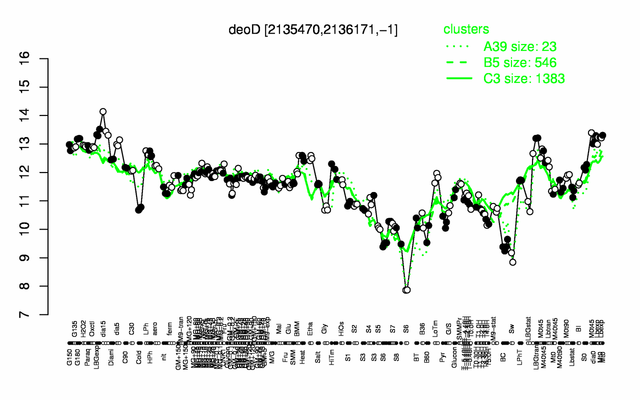
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of nucleotides, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU19630
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Purine nucleoside + phosphate = purine + alpha-D-ribose 1-phosphate (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- forms hexamers PubMed
- Localization:
- membrane (according to Swiss-Prot)
Database entries
- UniProt: O34925
- KEGG entry: [2]
- E.C. number: 2.4.2.1
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Priscila O de Giuseppe, Nadia H Martins, Andreia N Meza, Camila R dos Santos, Humberto D'Muniz Pereira, Mario T Murakami
Insights into phosphate cooperativity and influence of substrate modifications on binding and catalysis of hexameric purine nucleoside phosphorylases.
PLoS One: 2012, 7(9);e44282
[PubMed:22957058]
[WorldCat.org]
[DOI]
(I p)
Haojian Li, Guoqiang Zhang, Aihua Deng, Ning Chen, Tingyi Wen
De novo engineering and metabolic flux analysis of inosine biosynthesis in Bacillus subtilis.
Biotechnol Lett: 2011, 33(8);1575-80
[PubMed:21424839]
[WorldCat.org]
[DOI]
(I p)
Rosa Grenha, Vladimir M Levdikov, Mark J Fogg, Elena V Blagova, James A Brannigan, Anthony J Wilkinson, Keith S Wilson
Structure of purine nucleoside phosphorylase (DeoD) from Bacillus anthracis.
Acta Crystallogr Sect F Struct Biol Cryst Commun: 2005, 61(Pt 5);459-62
[PubMed:16511068]
[WorldCat.org]
[DOI]
(I p)