Difference between revisions of "CtsR"
(→Extended information on the protein) |
|||
| Line 72: | Line 72: | ||
* '''Effectors of protein activity:''' probably activated by dephosphorylation by [[McsA]] and inactivated by phosphorylation by [[McsB]] [http://www.ncbi.nlm.nih.gov/sites/entrez/14984053 PubMed1] [http://www.ncbi.nlm.nih.gov/sites/entrez/19498169 PubMed2], regulated proteolysis by [[ClpP]]/[[ClpC]] [http://www.ncbi.nlm.nih.gov/pubmed/11179229 PubMed], [http://www.ncbi.nlm.nih.gov/pubmed/16163393 PubMed], [http://www.ncbi.nlm.nih.gov/pubmed/17380125 PubMed] | * '''Effectors of protein activity:''' probably activated by dephosphorylation by [[McsA]] and inactivated by phosphorylation by [[McsB]] [http://www.ncbi.nlm.nih.gov/sites/entrez/14984053 PubMed1] [http://www.ncbi.nlm.nih.gov/sites/entrez/19498169 PubMed2], regulated proteolysis by [[ClpP]]/[[ClpC]] [http://www.ncbi.nlm.nih.gov/pubmed/11179229 PubMed], [http://www.ncbi.nlm.nih.gov/pubmed/16163393 PubMed], [http://www.ncbi.nlm.nih.gov/pubmed/17380125 PubMed] | ||
| − | * '''Interactions:''' [[McsA]]-[[ | + | * '''Interactions:''' [[McsA]]-[[McsB]]-[[CtsR]] |
* '''Localization:''' | * '''Localization:''' | ||
Revision as of 13:09, 20 January 2010
| Gene name | ctsR |
| Synonyms | yacG |
| Essential | no |
| Product | transcription repressor |
| Function | regulation of protein degradation |
| Regulatory function and regulation of this protein in SubtiPathways: Stress | |
| MW, pI | 17 kDa, 9.261 |
| Gene length, protein length | 462 bp, 154 aa |
| Immediate neighbours | rrnW-5S, mcsA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 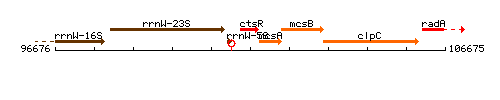
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Locus tag: BSU00830
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family: ctsR family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification: phosphorylation of a tyrosine residue by McsB PubMed, recently, it was reported thatCtsR is phosphorylatedby McsB on Arg-62 rather than on a tyrosine residue PubMed
- Cofactor(s):
- Effectors of protein activity: probably activated by dephosphorylation by McsA and inactivated by phosphorylation by McsB PubMed1 PubMed2, regulated proteolysis by ClpP/ClpC PubMed, PubMed, PubMed
- Localization:
Database entries
- Structure:
- UniProt: P37568
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Additional information: the mRNA is very stable (half-life > 15 min) PubMed
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Jakob Fuhrmann, Andreas Schmidt, Silvia Spiess, Anita Lehner, Kürsad Turgay, Karl Mechtler, Emmanuelle Charpentier, Tim Clausen
McsB is a protein arginine kinase that phosphorylates and inhibits the heat-shock regulator CtsR.
Science: 2009, 324(5932);1323-7
[PubMed:19498169]
[WorldCat.org]
[DOI]
(I p)
Janine Kirstein, David A Dougan, Ulf Gerth, Michael Hecker, Kürşad Turgay
The tyrosine kinase McsB is a regulated adaptor protein for ClpCP.
EMBO J: 2007, 26(8);2061-70
[PubMed:17380125]
[WorldCat.org]
[DOI]
(P p)
Marcus Miethke, Michael Hecker, Ulf Gerth
Involvement of Bacillus subtilis ClpE in CtsR degradation and protein quality control.
J Bacteriol: 2006, 188(13);4610-9
[PubMed:16788169]
[WorldCat.org]
[DOI]
(P p)
Janine Kirstein, Daniela Zühlke, Ulf Gerth, Kürşad Turgay, Michael Hecker
A tyrosine kinase and its activator control the activity of the CtsR heat shock repressor in B. subtilis.
EMBO J: 2005, 24(19);3435-45
[PubMed:16163393]
[WorldCat.org]
[DOI]
(P p)
Wolfgang Schumann
The Bacillus subtilis heat shock stimulon.
Cell Stress Chaperones: 2003, 8(3);207-17
[PubMed:14984053]
[WorldCat.org]
[DOI]
(P p)
Pekka Varmanen, Finn K Vogensen, Karin Hammer, Airi Palva, Hanne Ingmer
ClpE from Lactococcus lactis promotes repression of CtsR-dependent gene expression.
J Bacteriol: 2003, 185(17);5117-24
[PubMed:12923084]
[WorldCat.org]
[DOI]
(P p)
G Hambraeus, C von Wachenfeldt, L Hederstedt
Genome-wide survey of mRNA half-lives in Bacillus subtilis identifies extremely stable mRNAs.
Mol Genet Genomics: 2003, 269(5);706-14
[PubMed:12884008]
[WorldCat.org]
[DOI]
(P p)
J D Helmann, M F Wu, P A Kobel, F J Gamo, M Wilson, M M Morshedi, M Navre, C Paddon
Global transcriptional response of Bacillus subtilis to heat shock.
J Bacteriol: 2001, 183(24);7318-28
[PubMed:11717291]
[WorldCat.org]
[DOI]
(P p)
A Petersohn, M Brigulla, S Haas, J D Hoheisel, U Völker, M Hecker
Global analysis of the general stress response of Bacillus subtilis.
J Bacteriol: 2001, 183(19);5617-31
[PubMed:11544224]
[WorldCat.org]
[DOI]
(P p)
E Krüger, D Zühlke, E Witt, H Ludwig, M Hecker
Clp-mediated proteolysis in Gram-positive bacteria is autoregulated by the stability of a repressor.
EMBO J: 2001, 20(4);852-63
[PubMed:11179229]
[WorldCat.org]
[DOI]
(P p)
I Derré, G Rapoport, T Msadek
The CtsR regulator of stress response is active as a dimer and specifically degraded in vivo at 37 degrees C.
Mol Microbiol: 2000, 38(2);335-47
[PubMed:11069659]
[WorldCat.org]
[DOI]
(P p)
I Derré, G Rapoport, T Msadek
CtsR, a novel regulator of stress and heat shock response, controls clp and molecular chaperone gene expression in gram-positive bacteria.
Mol Microbiol: 1999, 31(1);117-31
[PubMed:9987115]
[WorldCat.org]
[DOI]
(P p)
E Krüger, M Hecker
The first gene of the Bacillus subtilis clpC operon, ctsR, encodes a negative regulator of its own operon and other class III heat shock genes.
J Bacteriol: 1998, 180(24);6681-8
[PubMed:9852015]
[WorldCat.org]
[DOI]
(P p)
E Krüger, T Msadek, M Hecker
Alternate promoters direct stress-induced transcription of the Bacillus subtilis clpC operon.
Mol Microbiol: 1996, 20(4);713-23
[PubMed:8793870]
[WorldCat.org]
[DOI]
(P p)
E Krüger, U Völker, M Hecker
Stress induction of clpC in Bacillus subtilis and its involvement in stress tolerance.
J Bacteriol: 1994, 176(11);3360-7
[PubMed:8195092]
[WorldCat.org]
[DOI]
(P p)