CtaC
- Description: cytochrome-c oxidase (subunit II)
| Gene name | ctaC |
| Synonyms | |
| Essential | no |
| Product | cytochrome-c oxidase (subunit II) |
| Function | respiration |
| Gene expression levels in SubtiExpress: ctaC | |
| Interactions involving this protein in SubtInteract: CtaC | |
| MW, pI | 39 kDa, 8.635 |
| Gene length, protein length | 1068 bp, 356 aa |
| Immediate neighbours | ctaB, ctaD |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 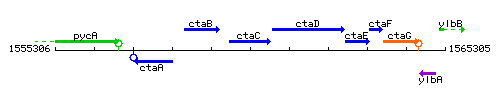
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed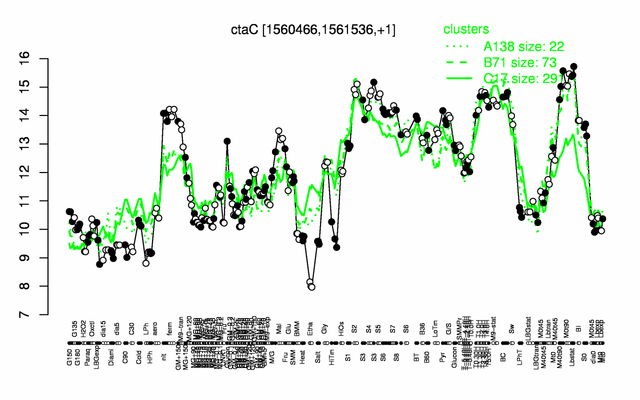
| |
Contents
Categories containing this gene/protein
respiration, membrane proteins
This gene is a member of the following regulons
Abh regulon, AbrB regulon, CcpA regulon, ResD regulon
The gene
Basic information
- Locus tag: BSU14890
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: 4 ferrocytochrome c + O2 + 4 H+ = 4 ferricytochrome c + 2 H2O (according to Swiss-Prot)
- Protein family: cytochrome c oxidase subunit 2 family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization: cell membrane (according to Swiss-Prot)
Database entries
- Structure:
- UniProt: P24011
- KEGG entry: [3]
- E.C. number: 1.9.3.1
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Ilana Kolodkin-Gal, Alexander K W Elsholz, Christine Muth, Peter R Girguis, Roberto Kolter, Richard Losick
Respiration control of multicellularity in Bacillus subtilis by a complex of the cytochrome chain with a membrane-embedded histidine kinase.
Genes Dev: 2013, 27(8);887-99
[PubMed:23599347]
[WorldCat.org]
[DOI]
(I p)
Onuma Chumsakul, Hiroki Takahashi, Taku Oshima, Takahiro Hishimoto, Shigehiko Kanaya, Naotake Ogasawara, Shu Ishikawa
Genome-wide binding profiles of the Bacillus subtilis transition state regulator AbrB and its homolog Abh reveals their interactive role in transcriptional regulation.
Nucleic Acids Res: 2011, 39(2);414-28
[PubMed:20817675]
[WorldCat.org]
[DOI]
(I p)
Hans-Matti Blencke, Georg Homuth, Holger Ludwig, Ulrike Mäder, Michael Hecker, Jörg Stülke
Transcriptional profiling of gene expression in response to glucose in Bacillus subtilis: regulation of the central metabolic pathways.
Metab Eng: 2003, 5(2);133-49
[PubMed:12850135]
[WorldCat.org]
[DOI]
(P p)
N Azarkina, S Siletsky, V Borisov, C von Wachenfeldt, L Hederstedt, A A Konstantinov
A cytochrome bb'-type quinol oxidase in Bacillus subtilis strain 168.
J Biol Chem: 1999, 274(46);32810-7
[PubMed:10551842]
[WorldCat.org]
[DOI]
(P p)
J Bengtsson, H Tjalsma, C Rivolta, L Hederstedt
Subunit II of Bacillus subtilis cytochrome c oxidase is a lipoprotein.
J Bacteriol: 1999, 181(2);685-8
[PubMed:9882689]
[WorldCat.org]
[DOI]
(P p)
X Liu, H W Taber
Catabolite regulation of the Bacillus subtilis ctaBCDEF gene cluster.
J Bacteriol: 1998, 180(23);6154-63
[PubMed:9829923]
[WorldCat.org]
[DOI]
(P p)
J van der Oost, C von Wachenfeld, L Hederstedt, M Saraste
Bacillus subtilis cytochrome oxidase mutants: biochemical analysis and genetic evidence for two aa3-type oxidases.
Mol Microbiol: 1991, 5(8);2063-72
[PubMed:1685007]
[WorldCat.org]
[DOI]
(P p)