Difference between revisions of "CpgA"
(→References) |
|||
| Line 120: | Line 120: | ||
# Hunt, A., Rawlins, J. P., Thomaides, H. B., and Errington, J. (2006) Functional analysis of 11 putative essential genes in Bacillus subtilis. Microbiology 152, 2895-2907. [http://www.ncbi.nlm.nih.gov/sites/entrez/17005971 PubMed] | # Hunt, A., Rawlins, J. P., Thomaides, H. B., and Errington, J. (2006) Functional analysis of 11 putative essential genes in Bacillus subtilis. Microbiology 152, 2895-2907. [http://www.ncbi.nlm.nih.gov/sites/entrez/17005971 PubMed] | ||
# Absalon C, Obuchowski M, Madec E, Delattre D, Holland IB, Séror SJ (2009) CpgA, EF-Tu and the stressosome protein YezB are substrates of the Ser/Thr kinase/phosphatase couple, PrkC/PrpC, in ''Bacillus subtilis''. ''Microbiology'' '''155:''' 932-943. [http://www.ncbi.nlm.nih.gov/sites/entrez/19246764 PubMed] | # Absalon C, Obuchowski M, Madec E, Delattre D, Holland IB, Séror SJ (2009) CpgA, EF-Tu and the stressosome protein YezB are substrates of the Ser/Thr kinase/phosphatase couple, PrkC/PrpC, in ''Bacillus subtilis''. ''Microbiology'' '''155:''' 932-943. [http://www.ncbi.nlm.nih.gov/sites/entrez/19246764 PubMed] | ||
| − | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/ | + | # Absalon C, Hamze K, Blanot D, Frehel C, Carballido-Lopez R, Holland BI, van Heijenoort J, Séror SJ. (2008) The GTPase CpgA is implicated in the deposition of the peptidoglycan sacculus in Bacillus subtilis. ''J Bacteriol.'' '''May;190(10):''' 3786-90. [http://www.ncbi.nlm.nih.gov/sites/entrez/PMID PubMed] |
| + | # Author1, Author2 & Author3 (year) Title ''Journal'' '''volume:''' page-page. [http://www.ncbi.nlm.nih.gov/sites/entrez/18344364 PubMed] | ||
Revision as of 15:00, 17 March 2009
- Description: write here
| Gene name | cpgA |
| Synonyms | yloQ |
| Essential | no |
| Product | |
| Function | unknown |
| MW, pI | 33 kDa, 4.743 |
| Gene length, protein length | 894 bp, 298 aa |
| Immediate neighbours | prkC, rpe |
| Gene sequence (+200bp) | Protein sequence |
Genetic context 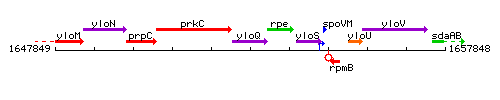
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity:
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Cofactor(s):
- Effectors of protein activity:
- Interactions:
- Localization:
Database entries
- Structure:
- Swiss prot entry:
- KEGG entry:
- E.C. number:
Additional information
Expression and regulation
- Operon:
- Sigma factor:
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
- Hunt, A., Rawlins, J. P., Thomaides, H. B., and Errington, J. (2006) Functional analysis of 11 putative essential genes in Bacillus subtilis. Microbiology 152, 2895-2907. PubMed
- Absalon C, Obuchowski M, Madec E, Delattre D, Holland IB, Séror SJ (2009) CpgA, EF-Tu and the stressosome protein YezB are substrates of the Ser/Thr kinase/phosphatase couple, PrkC/PrpC, in Bacillus subtilis. Microbiology 155: 932-943. PubMed
- Absalon C, Hamze K, Blanot D, Frehel C, Carballido-Lopez R, Holland BI, van Heijenoort J, Séror SJ. (2008) The GTPase CpgA is implicated in the deposition of the peptidoglycan sacculus in Bacillus subtilis. J Bacteriol. May;190(10): 3786-90. PubMed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed