Difference between revisions of "ComGA"
| Line 139: | Line 139: | ||
<pubmed> 15083159 </pubmed> | <pubmed> 15083159 </pubmed> | ||
==Original publications== | ==Original publications== | ||
| − | <pubmed>11814663,9422590,16009133,11298275,17630974,9723928,21278288,12107147, 8196543 2507524 21564336</pubmed> | + | <pubmed>11814663,9422590,16009133,11298275,17630974,9723928,21278288,12107147, 8196543 2507524 21564336 21707789 </pubmed> |
[[Category:Protein-coding genes]] | [[Category:Protein-coding genes]] | ||
Revision as of 11:07, 29 June 2011
- Description: late competence protein, traffic ATPase, binds and transports transforming DNA
| Gene name | comGA |
| Synonyms | |
| Essential | no |
| Product | traffic ATPase |
| Function | genetic competence |
| Metabolic function and regulation of this protein in SubtiPathways: Protein secretion | |
| MW, pI | 40 kDa, 8.893 |
| Gene length, protein length | 1068 bp, 356 aa |
| Immediate neighbours | comGB, corA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 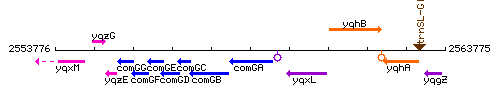
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
genetic competence, membrane proteins
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU24730
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: binds and transports transforming DNA
- Protein family: GSP E family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Localization:
Database entries
- Structure:
- UniProt: P25953
- KEGG entry: [3]
- E.C. number:
Additional information
Expression and regulation
- Regulation:
- repressed by casamino acids PubMed
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Inês Chen, David Dubnau
DNA uptake during bacterial transformation.
Nat Rev Microbiol: 2004, 2(3);241-9
[PubMed:15083159]
[WorldCat.org]
[DOI]
(P p)
Original publications
Kenneth Briley, Angella Dorsey-Oresto, Peter Prepiak, Miguel J Dias, Jessica M Mann, David Dubnau
The secretion ATPase ComGA is required for the binding and transport of transforming DNA.
Mol Microbiol: 2011, 81(3);818-30
[PubMed:21707789]
[WorldCat.org]
[DOI]
(I p)
Kenneth Briley, Peter Prepiak, Miguel J Dias, Jeanette Hahn, David Dubnau
Maf acts downstream of ComGA to arrest cell division in competent cells of B. subtilis.
Mol Microbiol: 2011, 81(1);23-39
[PubMed:21564336]
[WorldCat.org]
[DOI]
(I p)
Miriam Kaufenstein, Martin van der Laan, Peter L Graumann
The three-layered DNA uptake machinery at the cell pole in competent Bacillus subtilis cells is a stable complex.
J Bacteriol: 2011, 193(7);1633-42
[PubMed:21278288]
[WorldCat.org]
[DOI]
(I p)
Naomi Kramer, Jeanette Hahn, David Dubnau
Multiple interactions among the competence proteins of Bacillus subtilis.
Mol Microbiol: 2007, 65(2);454-64
[PubMed:17630974]
[WorldCat.org]
[DOI]
(P p)
Jeanette Hahn, Berenike Maier, Bert Jan Haijema, Michael Sheetz, David Dubnau
Transformation proteins and DNA uptake localize to the cell poles in Bacillus subtilis.
Cell: 2005, 122(1);59-71
[PubMed:16009133]
[WorldCat.org]
[DOI]
(P p)
Ulrike Mäder, Georg Homuth, Christian Scharf, Knut Büttner, Rüdiger Bode, Michael Hecker
Transcriptome and proteome analysis of Bacillus subtilis gene expression modulated by amino acid availability.
J Bacteriol: 2002, 184(15);4288-95
[PubMed:12107147]
[WorldCat.org]
[DOI]
(P p)
Hanne Jarmer, Randy Berka, Steen Knudsen, Hans H Saxild
Transcriptome analysis documents induced competence of Bacillus subtilis during nitrogen limiting conditions.
FEMS Microbiol Lett: 2002, 206(2);197-200
[PubMed:11814663]
[WorldCat.org]
[DOI]
(P p)
B J Haijema, J Hahn, J Haynes, D Dubnau
A ComGA-dependent checkpoint limits growth during the escape from competence.
Mol Microbiol: 2001, 40(1);52-64
[PubMed:11298275]
[WorldCat.org]
[DOI]
(P p)
Y S Chung, F Breidt, D Dubnau
Cell surface localization and processing of the ComG proteins, required for DNA binding during transformation of Bacillus subtilis.
Mol Microbiol: 1998, 29(3);905-13
[PubMed:9723928]
[WorldCat.org]
[DOI]
(P p)
Y S Chung, D Dubnau
All seven comG open reading frames are required for DNA binding during transformation of competent Bacillus subtilis.
J Bacteriol: 1998, 180(1);41-5
[PubMed:9422590]
[WorldCat.org]
[DOI]
(P p)
D van Sinderen, A ten Berge, B J Hayema, L Hamoen, G Venema
Molecular cloning and sequence of comK, a gene required for genetic competence in Bacillus subtilis.
Mol Microbiol: 1994, 11(4);695-703
[PubMed:8196543]
[WorldCat.org]
[DOI]
(P p)
M Albano, R Breitling, D A Dubnau
Nucleotide sequence and genetic organization of the Bacillus subtilis comG operon.
J Bacteriol: 1989, 171(10);5386-404
[PubMed:2507524]
[WorldCat.org]
[DOI]
(P p)