Difference between revisions of "ClpC"
(→Expression and regulation) |
(→Biological materials) |
||
| Line 103: | Line 103: | ||
=Biological materials = | =Biological materials = | ||
| − | * '''Mutant:''' ''clpC::tet'' available from the | + | * '''Mutant:''' ''clpC::tet'' available from the [http://subtiwiki.uni-goettingen.de/wiki/index.php/Leendert_Hamoen Hamoen]] Lab |
* '''Expression vector:''' | * '''Expression vector:''' | ||
| Line 109: | Line 109: | ||
* '''lacZ fusion:''' | * '''lacZ fusion:''' | ||
| − | * '''GFP fusion:''' C-terminal GFP fusions (single copy, also as CFP and YFP variants) available from the | + | * '''GFP fusion:''' C-terminal GFP fusions (single copy, also as CFP and YFP variants) available from the [http://subtiwiki.uni-goettingen.de/wiki/index.php/Leendert_Hamoen Hamoen]] Lab |
* '''two-hybrid system:''' | * '''two-hybrid system:''' | ||
Revision as of 21:57, 28 April 2009
- Description: ATP-dependent Clp protease, ATPase subunit
| Gene name | clpC |
| Synonyms | mecB |
| Essential | no |
| Product | ATP-dependent Clp protease, ATPase subunit |
| Function | protein degradation
positive regulator of autolysin (LytC and LytD) synthesis |
| MW, pI | 89 kDa, 5.746 |
| Gene length, protein length | 2430 bp, 810 aa |
| Immediate neighbours | mcsB, radA |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 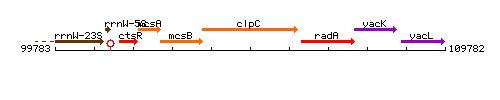
This image was kindly provided by SubtiList
| |
Contents
The gene
Basic information
- Coordinates:
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATPase/chaperone
Extended information on the protein
- Kinetic information:
- Domains: AAA-ATPase PFAM
- Modification:
- Cofactor(s):
- Effectors of protein activity:
- Interactions: YpbH-ClpC, McsB-ClpC, ClpC-MecA, ClpC-ClpP, YpbH-ClpC, McsB-ClpC, ClpC-MecA, ClpC-ClpP, YpbH-ClpC, McsB-ClpC, ClpC-MecA, ClpC-ClpP, YpbH-ClpC, McsB-ClpC, ClpC-MecA, ClpC-ClpP
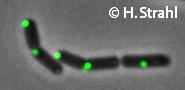
- Localization: cytoplasmic polar clusters, excluded from the nucleoid, induced clustering upon heatshock, colocalization with ClpP Pubmed
Database entries
- Structure:
- Swiss prot entry: P37571
- KEGG entry: [3]
- E.C. number:
Additional information
- subject to Clp-dependent proteolysis upon glucose starvation PubMed
Expression and regulation
- Operon:
- Additional information: subject to Clp-dependent proteolysis upon glucose starvation PubMed
Biological materials
- Mutant: clpC::tet available from the Hamoen] Lab
- Expression vector:
- lacZ fusion:
- GFP fusion: C-terminal GFP fusions (single copy, also as CFP and YFP variants) available from the Hamoen] Lab
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Leendert Hamoen, Newcastle University, UK homepage
Your additional remarks
References
- Petersohn et al. (2001) Global Analysis of the General Stress Response of Bacillus subtilis. J Bacteriol. 183: 5617-5631 PubMed
- Gerth et al. (2008) Clp-dependent proteolysis down-regulates central metabolic pathways in glucose-starved Bacillus subtilis. J Bacteriol 190:321-331 PubMed
- Kock H, Gerth U, Hecker M. (2004) MurAA, catalysing the first committed step in peptidoglycan biosynthesis, is a target of Clp-dependent proteolysis in Bacillus subtilis. Mol Microbiol, 51:1087-1102. PubMed
- Kirstein, J., Strahl, H., Molière, N., Hamoen, LW., Turgay K. (2008) Localization of general and regulatory proteolysis in Bacillus subtilis cells. Mol Microbiol. 70:682-94. Pubmed
- Author1, Author2 & Author3 (year) Title Journal volume: page-page. PubMed