Difference between revisions of "Bcd"
(→Expression and regulation) |
|||
| Line 117: | Line 117: | ||
* '''Regulatory mechanism:''' | * '''Regulatory mechanism:''' | ||
| − | ** [[CodY]]: | + | ** [[CodY]]: indirect regulation {{PubMed|24843172}} |
** [[BkdR]]: transcription activation {{PubMed|10094682}} | ** [[BkdR]]: transcription activation {{PubMed|10094682}} | ||
Revision as of 11:24, 21 May 2014
- Description: valine dehydrogenase, isoleucine dehydrogenase, L-leucine dehydrogenase
| Gene name | bcd |
| Synonyms | bkd, yqiT |
| Essential | no |
| Product | valine dehydrogenase, isoleucine dehydrogenase, L-leucine dehydrogenase |
| Function | utilization of branched-chain keto acids |
| Gene expression levels in SubtiExpress: bcd | |
| Metabolic function and regulation of this protein in SubtiPathways: bcd | |
| MW, pI | 39 kDa, 4.942 |
| Gene length, protein length | 1092 bp, 364 aa |
| Immediate neighbours | buk, ptb |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 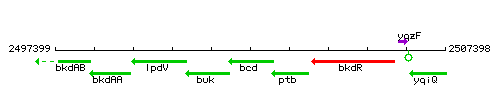
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed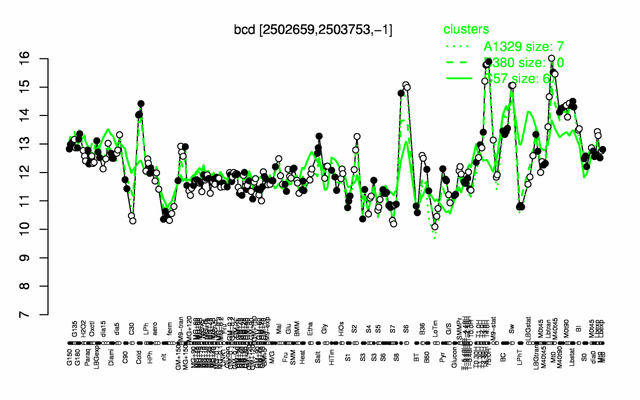
| |
Contents
Categories containing this gene/protein
This gene is a member of the following regulons
BkdR regulon, CodY regulon, SigL regulon
The gene
Basic information
- Locus tag: BSU24080
Phenotypes of a mutant
Database entries
- BsubCyc: BSU24080
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: L-leucine + H2O + NAD+ = 4-methyl-2-oxopentanoate + NH3 + NADH (according to Swiss-Prot)
- Protein family: Glu/Leu/Phe/Val dehydrogenases family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- BsubCyc: BSU24080
- UniProt: P54531
- KEGG entry: [3]
- E.C. number: 1.4.1.9
Additional information
Expression and regulation
- Additional information:
- number of protein molecules per cell (minimal medium with glucose and ammonium, exponential phase): 3408 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, early stationary phase after glucose exhaustion): 2690 PubMed
- number of protein molecules per cell (minimal medium with glucose and ammonium, late stationary phase after glucose exhaustion): 11556 PubMed
Biological materials
- Mutant:
- a bcd mutant and a bcd ybgE ywaA triple mutant are available in Linc Sonenshein's lab
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Reviews
Original publications
M Nickel, G Homuth, C Böhnisch, U Mäder, T Schweder
Cold induction of the Bacillus subtilis bkd operon is mediated by increased mRNA stability.
Mol Genet Genomics: 2004, 272(1);98-107
[PubMed:15241682]
[WorldCat.org]
[DOI]
(P p)
Ken-ichi Yoshida, Hirotake Yamaguchi, Masaki Kinehara, Yo-hei Ohki, Yoshiko Nakaura, Yasutaro Fujita
Identification of additional TnrA-regulated genes of Bacillus subtilis associated with a TnrA box.
Mol Microbiol: 2003, 49(1);157-65
[PubMed:12823818]
[WorldCat.org]
[DOI]
(P p)
Tanja Kaan, Georg Homuth, Ulrike Mäder, Julia Bandow, Thomas Schweder
Genome-wide transcriptional profiling of the Bacillus subtilis cold-shock response.
Microbiology (Reading): 2002, 148(Pt 11);3441-3455
[PubMed:12427936]
[WorldCat.org]
[DOI]
(P p)
M Debarbouille, R Gardan, M Arnaud, G Rapoport
Role of bkdR, a transcriptional activator of the sigL-dependent isoleucine and valine degradation pathway in Bacillus subtilis.
J Bacteriol: 1999, 181(7);2059-66
[PubMed:10094682]
[WorldCat.org]
[DOI]
(P p)