Difference between revisions of "BacD"
(→References) |
(→Database entries) |
||
| Line 84: | Line 84: | ||
=== Database entries === | === Database entries === | ||
| − | * '''Structure:''' | + | * '''Structure:''' [http://www.rcsb.org/pdb/cgi/explore.cgi?pdbId=3VMM 3VMM] |
* '''UniProt:''' [http://www.uniprot.org/uniprot/P39641 P39641] | * '''UniProt:''' [http://www.uniprot.org/uniprot/P39641 P39641] | ||
Revision as of 09:15, 28 December 2011
- Description: alanine-anticapsin ligase
| Gene name | bacD |
| Synonyms | ywfE, ipa-83d |
| Essential | no |
| Product | alanine-anticapsin ligase |
| Function | biosynthesis of the antibiotic bacilysin |
| MW, pI | 52 kDa, 4.679 |
| Gene length, protein length | 1416 bp, 472 aa |
| Immediate neighbours | bacE, bacC |
| Get the DNA and protein sequences (Barbe et al., 2009) | |
Genetic context 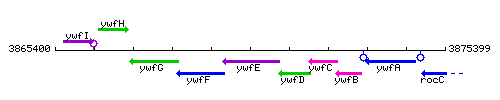
This image was kindly provided by SubtiList
| |
Contents
Categories containing this gene/protein
miscellaneous metabolic pathways, biosynthesis of antibacterial compounds
This gene is a member of the following regulons
AbrB regulon, CodY regulon, ScoC regulon
The gene
Basic information
- Locus tag: BSU37710
Phenotypes of a mutant
Database entries
- DBTBS entry: no entry
- SubtiList entry: [1]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: ATP + an L-amino acid + an L-amino acid = ADP + phosphate + L-aminoacyl-L-amino acid (according to Swiss-Prot)
- Protein family:
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 3VMM
- UniProt: P39641
- KEGG entry: [2]
- E.C. number: 6.3.2.28
Additional information
Expression and regulation
- Regulation:
- Regulatory mechanism:
- Additional information:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Additional publications: PubMed
Takashi Inaoka, Kozo Ochi
Scandium stimulates the production of amylase and bacilysin in Bacillus subtilis.
Appl Environ Microbiol: 2011, 77(22);8181-3
[PubMed:21948839]
[WorldCat.org]
[DOI]
(I p)
Takashi Inaoka, Guojun Wang, Kozo Ochi
ScoC regulates bacilysin production at the transcription level in Bacillus subtilis.
J Bacteriol: 2009, 191(23);7367-71
[PubMed:19801406]
[WorldCat.org]
[DOI]
(I p)
Kuniki Kino, Yoichi Kotanaka, Toshinobu Arai, Makoto Yagasaki
A novel L-amino acid ligase from Bacillus subtilis NBRC3134, a microorganism producing peptide-antibiotic rhizocticin.
Biosci Biotechnol Biochem: 2009, 73(4);901-7
[PubMed:19352016]
[WorldCat.org]
[DOI]
(I p)
Kazuhiko Tabata, Hajime Ikeda, Shin-Ichi Hashimoto
ywfE in Bacillus subtilis codes for a novel enzyme, L-amino acid ligase.
J Bacteriol: 2005, 187(15);5195-202
[PubMed:16030213]
[WorldCat.org]
[DOI]
(P p)
Takashi Inaoka, Kosaku Takahashi, Mayumi Ohnishi-Kameyama, Mitsuru Yoshida, Kozo Ochi
Guanine nucleotides guanosine 5'-diphosphate 3'-diphosphate and GTP co-operatively regulate the production of an antibiotic bacilysin in Bacillus subtilis.
J Biol Chem: 2003, 278(4);2169-76
[PubMed:12372825]
[WorldCat.org]
[DOI]
(P p)