AroE
- Description: 3-phosphoshikimate 1-carboxyvinyltransferase
| Gene name | aroE |
| Synonyms | |
| Essential | no |
| Product | 3-phosphoshikimate 1-carboxyvinyltransferase |
| Function | biosynthesis of aromatic amino acids |
| Gene expression levels in SubtiExpress: aroE | |
| Metabolic function and regulation of this protein in SubtiPathways: Phe, Tyr, Trp | |
| MW, pI | 45 kDa, 6.341 |
| Gene length, protein length | 1284 bp, 428 aa |
| Immediate neighbours | ypiA, tyrA |
| Sequences | Protein DNA DNA_with_flanks |
Genetic context 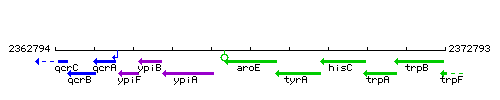
This image was kindly provided by SubtiList
| |
Expression at a glance PubMed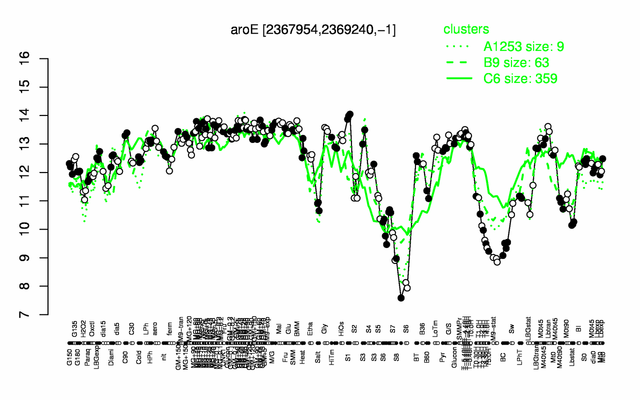
| |
Contents
Categories containing this gene/protein
biosynthesis/ acquisition of amino acids
This gene is a member of the following regulons
The gene
Basic information
- Locus tag: BSU22600
Phenotypes of a mutant
Database entries
- DBTBS entry: [1]
- SubtiList entry: [2]
Additional information
The protein
Basic information/ Evolution
- Catalyzed reaction/ biological activity: Phosphoenolpyruvate + 3-phosphoshikimate = phosphate + 5-O-(1-carboxyvinyl)-3-phosphoshikimate (according to Swiss-Prot)
- Protein family: ABC transporter family (according to Swiss-Prot)
- Paralogous protein(s):
Extended information on the protein
- Kinetic information:
- Domains:
- Modification:
- Cofactor(s):
- Effectors of protein activity:
Database entries
- Structure: 3RMT (from B. halodurans, 68% identity, 87% similarity)
- UniProt: P20691
- KEGG entry: [3]
- E.C. number: 2.5.1.19
Additional information
Expression and regulation
- Operon:
- Regulatory mechanism:
Biological materials
- Mutant:
- Expression vector:
- lacZ fusion:
- GFP fusion:
- two-hybrid system:
- Antibody:
Labs working on this gene/protein
Your additional remarks
References
Martin Lehnik-Habrink, Marc Schaffer, Ulrike Mäder, Christine Diethmaier, Christina Herzberg, Jörg Stülke
RNA processing in Bacillus subtilis: identification of targets of the essential RNase Y.
Mol Microbiol: 2011, 81(6);1459-73
[PubMed:21815947]
[WorldCat.org]
[DOI]
(I p)
J Otridge, P Gollnick
MtrB from Bacillus subtilis binds specifically to trp leader RNA in a tryptophan-dependent manner.
Proc Natl Acad Sci U S A: 1993, 90(1);128-32
[PubMed:8419914]
[WorldCat.org]
[DOI]
(P p)
P Babitzke, P Gollnick, C Yanofsky
The mtrAB operon of Bacillus subtilis encodes GTP cyclohydrolase I (MtrA), an enzyme involved in folic acid biosynthesis, and MtrB, a regulator of tryptophan biosynthesis.
J Bacteriol: 1992, 174(7);2059-64
[PubMed:1551827]
[WorldCat.org]
[DOI]
(P p)
D J Henner, L Band, G Flaggs, E Chen
The organization and nucleotide sequence of the Bacillus subtilis hisH, tyrA and aroE genes.
Gene: 1986, 49(1);147-52
[PubMed:3106153]
[WorldCat.org]
[DOI]
(P p)
D J Henner, L Band, H Shimotsu
Nucleotide sequence of the Bacillus subtilis tryptophan operon.
Gene: 1985, 34(2-3);169-77
[PubMed:3924737]
[WorldCat.org]
[DOI]
(P p)
H Shimotsu, D J Henner
Characterization of the Bacillus subtilis tryptophan promoter region.
Proc Natl Acad Sci U S A: 1984, 81(20);6315-9
[PubMed:6436812]
[WorldCat.org]
[DOI]
(P p)